From Surf Wiki (app.surf) — the open knowledge base
Protein trimer
Macromolecular complex formed by three macromolecules
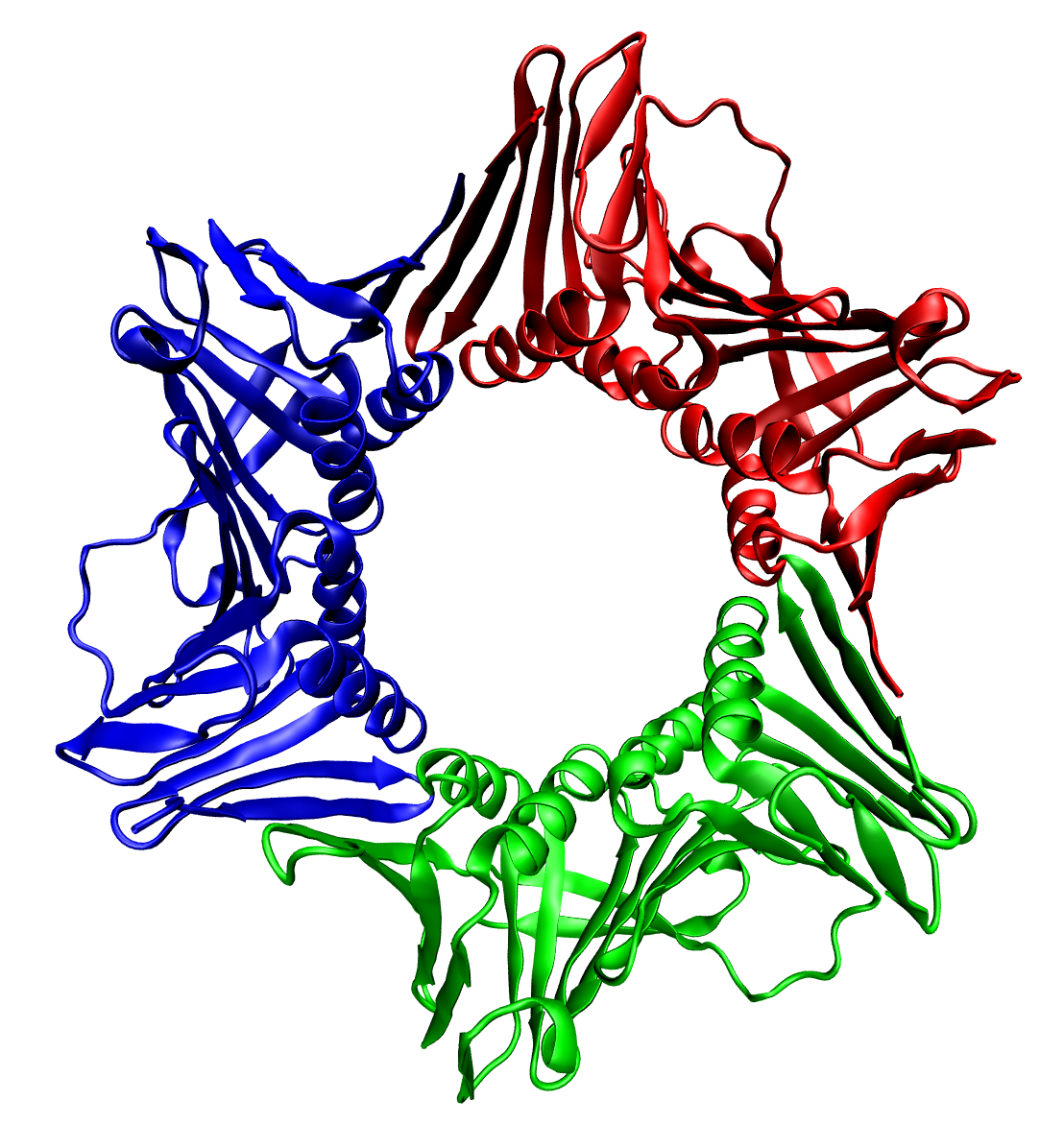
Macromolecular complex formed by three macromolecules

thumbnail|right|400px|Trimeric form of a TNF-α mutant In biochemistry, a protein trimer is a macromolecular complex formed by three, usually non-covalently bound, macromolecules like proteins or nucleic acids. A protein trimer often occurs from the assembly of a protein's quaternary structure. The non-covalent interactions between the hydrophobic and hydrophilic regions on the polypeptides units help to stabilize the quaternary structure. Since a protein trimer is composed of multiple polypeptide subunits, it is considered an oligomer.
A homotrimer would be formed by three identical molecules. A heterotrimer would be formed by three different macromolecules. Type II Collagen is an example of homotrimeric protein, while Type I collagen is an AAB-type heterotrimeric protein. An example of viral protein homotrimeric protein is mammarenavirus of Z matrix protein.
Porins usually arrange themselves in membranes as trimers.
TOC
Bacteriophage T4 tail fiber
Multiple copies of a polypeptide encoded by a gene often can form an aggregate referred to as a multimer. When a multimer is formed from polypeptides produced by two different mutant alleles of a particular gene, the mixed multimer may exhibit greater functional activity than the unmixed multimers formed by each of the mutants alone. When a mixed multimer displays increased functionality relative to the unmixed multimers, the phenomenon is referred to as intragenic complementation. The distal portion of each of the bacteriophage T4 tail fibers is encoded by gene 37 and mutants defective in this gene undergo intragenic complementation. This finding indicated that the distal tail fibers are a multimer of the gene 37 encoded polypeptide. An analysis of the complementation data further indicated that the polypeptides making up the multimer were folded back on themselves in the form of a hairpin. A further high-resolution crystal structure analysis of the distal tail fiber indicated that the gene 37 polypeptides are present as a trimer and that each polypeptide of the trimer is folded back on itself in a hairpin configuration.
References
References
- Godbey, W.T.. (2014). "Proteins". Elsevier.
- Pelley, John W.. (2012-01-01). "3 - Protein Structure and Function". W.B. Saunders.
- Cha, S.S.. (1998). "High resolution crystal structure of a human tumor necrosis factor-alpha mutant with low systemic toxicity". J. Biol. Chem..
- (2025-04-29). "Mammarenavirus Z Protein Myristoylation and Oligomerization Are Not Required for Its Dose-Dependent Inhibitory Effect on vRNP Activity". BioChem.
- (Jan 1964). "The theory of inter-allelic complementation". J Mol Biol.
- (1965). "Intragenic complementation among temperature sensitive mutants of bacteriophage T4D". Genetics.
- (2010). "Structure of the bacteriophage T4 long tail fiber receptor-binding tip". Proc Natl Acad Sci U S A.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Protein trimer — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report