From Surf Wiki (app.surf) — the open knowledge base
PBDC1
Human gene

Human gene
CXorf26 (Chromosome X Open Reading Frame 26), also known as MGC874, is a well conserved human gene found on the plus strand of the short arm of the X chromosome. The exact function of the gene is poorly understood, but the polysaccharide biosynthesis domain that spans a major portion of the protein product (known as UPF0368), as well as the yeast homolog, YPL225, offer insights into its possible function.
Proposed function
Given the mass of data available on CXorf26, potential function is likely related to the workings of RNA polymerase II, ubiquitination, and ribosomes in the cytoplasm. The basis of these arguments is on the interaction data of human CXorf26 as well as its yeast homolog, YPL225W. Both homologs show interaction with multiple ubiquinated proteins as well as the transcriptional enzyme RNA polymerase II. For example, ubiquitiation and subsequent degradation of the 26S proteasome serves an important function in regulating transcription in eukaryotes. The yeast protein RPN11, which interacts with YPL225W, has a homolog in humans that is a metalloprotease component of 26S proteasome that also degrades proteins targeted for destruction by the ubiquitin pathway. These functions do not seem to relate to a polysaccharide biosynthesis function (as would be assumed due to its conserved domain), but it may still play a role in the secondary structure or sites of phosphorylation.
Further experimentation into the potential role of CXorf26 may give further insight into its exact function in these key cellular processes. Experiments such as using a RNA polymerase II inhibitor and subsequent gene expression of CXorf26 could reveal potential functions as well as examination of a complete knockout of YPL225W in yeast using methods such as RNAi.
Gene

CXorf26 is found on the plus strand of the short arm of the X chromosome, specifically on the gene locus Xq13.3 spanning the genomic chromosome region from bases 75,393,420-75,397,740. The primary mRNA transcript sequence has 1214 base pairs and its protein product, UPF0368, is composed of 233 amino acids and has a predicted mass of 26,057 Da. The locus where CXorf26 is located, Xq13.3, has known associations to X-linked mental retardation. The third gene located upstream of CXorf26 is ATRX, which encodes for an ATPase/helicase domain, and when mutated causes an X-linked mental retardation syndrome along with alpha thalassemia syndrome; both are known to cause changes in the DNA methylation patterns. Furthermore, the third gene downstream of CXorf26, ZDHHC15, which when mutated, causes mental retardation X-linked type 91. One noteworthy gene located nearby is Xist, which plays a role in the inactivation process of the X chromosome. X inactivation relates to CXorf26, and is discussed below in the relevant research section.
Expression
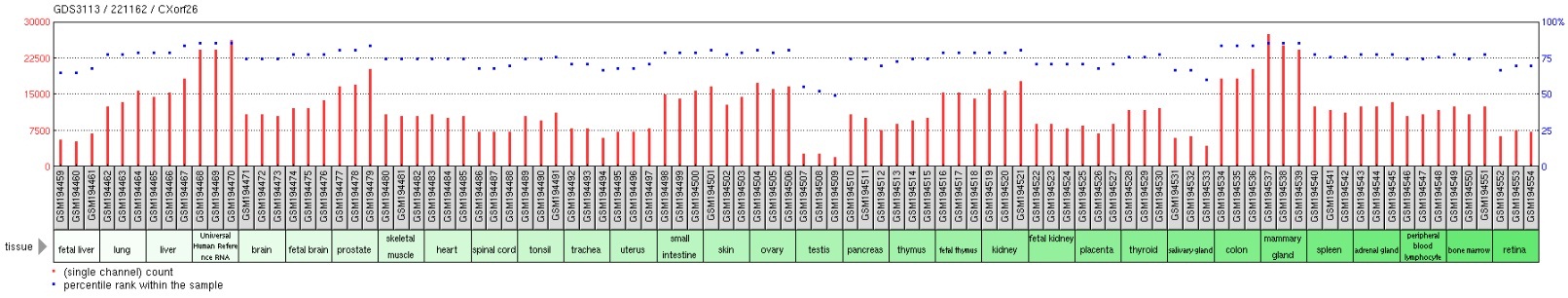

Gene expression profiles in the Gene Expression Omnibus (GEO) repository located within the NCBI website demonstrated that there were not many treatments that resulted in a changing of expression of CXorf26 in examined tissues. However, one experiment compared CXorf26 expression in lung adenocarcinoma CL1-5 cells either overexpressing or underexpressing Claudin-1. Results indicated that CXorf26 expression greatly drops when CLDN1 is overexpressed. CLDN1 is a major component in forming tight junction complexes between cells, which foster cell-cell adhesion of cell membranes. More tight junctions formed by CLDN1 would likely result in decreased expression of CXorf26 since the cell membrane would be used for tight junctions instead of its normal function related to heparan sulfate.
Alternative splice forms
There is only one alternative splice form for CXorf26. This splice form has significantly fewer mRNA base pairs at 977, but still has a protein product of 232 amino acids. This alternative splice form appears to be missing exon 5 of the transcript, but it may be added onto exon 6, creating a larger exon compared to the consensus transcript.
There were no other predicted exons within the genomic CXorf26 sequence when 3000 base pairs were added on either side in the search.
Promoter region
The promoter for CXorf26 is predicted to be located from bases 75392235 to 75393075 on the X chromosome positive strand. The promoter region has extensive conservation with all primates and most mammal homologs, but conservation is lessened in more distantly related species. Given the primary transcript begins at base 7539277, the promoter overlaps with it by 304 bases. 20 predicted transcription factor binding sites with their transcription factor family was collected as well. A high amount of the transcriptional factors relate to zinc finger factors, which have the function of stabilizing protein folds, while none of the factors seem to relate to a potential polysaccharide biosynthesis function. One transcription factor family predicted to bind to the promoter region was V$CHRF, and is involved in regulation of the cell cycle. The regulation could be related to ubiquitin function; proteins with ubiquitination type function were found to interact with CXorf26.
Protein
Subcellular distribution
The CXorf26 protein is 56.5% likely to be localized within the cytoplasm while 17.4% likely to localized to the mitochondria. CXorf26's yeast homolog, YPL225W, was GFP tagged and its location was determined to be in the cytoplasm. Cytoplasmic location instead of transmembrane was supported since no hydrophobic signal peptide sequence and TMAP predicted no potential transmembrane segments in CXorf26 or any of its homologs in other species.
Polysaccharide domain
CXorf26 was found to have conserved domain known as DUF757 within its sequence. The conserved domain spans a majority of the protein sequence, from amino acids 39-159. Conservation of the domain is strong throughout all homologs compared, including mammals, invertebrates such as insects, and even sponges. The yeast homolog, YPL225W, shows 42.4% identity and 62% similarity in this domain. Conservation of the domain is especially high in areas which include one of the multiple alpha helices or beta sheets. There are also multiple conserved phosphorylation sites located in the amino acid sequence at tyrosine 72 and serine 126.
According to NCBI, this domain is in the family of proteins expected play a role in xylan biosynthesis in plant cell walls, but its exact role in the synthesis pathway is unknown. As animal cells do not contain cell walls, its exact function in other organisms such as humans is unknown.
Xylan is made from units of the pentose sugar xylose, which is known for being the first saccharide in multiple biosynthetic pathways of anionic polysaccharides such as heparan sulfate and chondroitin sulfate. Like Xylan, heparan sulfate it is found on the cell surface; since it is needed for both the cell surface and extracellular matrix, it may explain CXorf26's high expression in nearly all human tissues. Heparan biosynthesis occurs in the lumen of the endoplasmic reticulum and is initiated by the transfer of a xylose from UDP-xylose by xylosyltransferase to specific serine residues within the protein core. PSORTII predicts the presence of a KKXX-like motif, GEKA, near the C-terminus of CXorf26. KKXX-like motifs are predicted endoplasmic reticulum membrane retention signals. This motif is only conserved in primates. However, another KKXX-like motif, QDKE, is found to exist at the end of the domain. The K in this motif is highly conserved back to most invertebrates. However, contradicting results from NetNGlyc predicted no N-glycosylation sites, suggesting CXorf26 does not undergo special folding in the endoplasmic reticulum lumen. Given that the conserved domain cannot function to create xylan since there are no cell walls in animal cells, the function may be related to this pathway.
Secondary structure
Predictions across multiple programs suggest the presence of 7 alpha helices and 2 beta sheets for CXorf26; the majority of the secondary structures are in the conserved domain. Experimental evidence in the yeast homolog shows 4 alpha helices and 2 beta sheets all in the polysaccharide domain, just as the predicted SWISS model above shows for humans. The location of the secondary structures are also conserved.
Post-translational modifications
Pepsin (pH 1.3), Asp-N endopeptidase, N-terminal Glutamate and Proteinase K all had 50 or more cleavage sites within the protein, but none of the 10 caspases had any cleavage sites. This suggests CXorf26 is not likely to be cleaved or degraded during apoptosis. This follows with the observation that CXorf26 is expressed highly in nearly all tissues and experimental conditions.
Lysine 63 and 66 are potential sites of glycation of epsilon amino groups of lysines. Lysine 63 was conserved in both Macaca mulatta and Bombus impatiens. There are 10 serine, 3 threonine, and 6 tyrosine phosphorylation sites predicted within the CXorf26 protein. When comparing the predicted phosphorylation sites, those shown in the table below were those conserved in Macaca mulatta as well as Bombus impatiens. S127 was left in the table even though Homo sapiens and Macaca mulatta did not have significant scores above threshold for that position. Through evolutionary change, the serine in Bombus was changed to a tyrosine in Homo sapiens and Macaca mulatta, which is still capable of phosphorylation, suggesting although there was a mutation, it would likely not result in a large change for the protein and its function.
| Bombus impatiens | Homo sapiens & Macaca mulatta |
|---|---|
| Serine 20 | Serine 23 |
| Serine 91 | Serine 94 |
| Tyrosine 69 | Tyrosine 72 |
| Tyrosine126 | Tyrosine 129 |
| *Serine 127** | *Tyrosine 130** |
Species distribution
CXorf26 is strongly evolutionary conserved, with conservation found in Batrachochytrium dendrobatidis. A multiple sequence alignment of 20 orthologous protein sequences reveals very strong conservation of the polysaccharide biosynthesis domain, but conservation after it was essentially non-existent in invertebrates. For those vertebrates that contained a sequence after the conserved domain, it was found to be of low complexity and filled with repetitive sequence of the amino acid motif 'GEK', corresponding to amino acids glycine, glutamic acid, and lysine. Glutamic acid and lysine both are charged, which contributes to the overall hydrophilicity of the section after the conserved domain.
| Species | Common name | Accession number | Length | Protein identity | Protein similarity |
|---|---|---|---|---|---|
| Homo sapiens | Human | NP_057584.2 | 233aa | 100% | 100% |
| Nomascus leucogenys | Gibbon | XP_003269034.1 | 233aa | 99% | 99% |
| Macaca mulatta | Rhesus monkey | NP_001181035.1 | 233aa | 98% | 98% |
| Callithrix jacchus | Marmoset | XP_002763066.1 | 232aa | 95% | 97% |
| Mus musculus | Mouse | NP_080588.1 | 198aa | 80% | 85% |
| Loxodonta africana | African elephant | XP_003412818.1 | 202aa | 80% | 88% |
| Ailuropoda melanoleuca | Giant panda | XP_002930750.1 | 219aa | 80% | 84% |
| Bos taurus | Cattle | XP_002700032.1 | 219aa | 78% | 86% |
| Monodelphis domestica | Opossum | XP_001381973.1 | 226aa | 59% | 89% |
| Oreochromis niloticus | Nile tilapia | XP_003453679.1 | 169aa | 46% | 83% |
| Bombus impatiens | Bumblebee | XP_003487356.1 | 168aa | 38% | 74% |
| Acromyrmex echinatior | Ant | EGI60293.1 | 197aa | 32% | 74% |
| Amphimedon queenslandica | Sponge | XP_003383281.1 | 159aa | 31% | 74% |
| Saccharomyces cerevisiae | Yeast | NP_015099.1 | 146aa | 27% | 62% |
| Batrachochytrium dendrobatidis | Fungus | EGF83065.1 | 74aa | 16% | 65% |
Yeast homolog YPL225W
The CXorf26 homolog in yeast, YPL225W, has an overall identity match of 27% but a 42.4% identity and 62% similarity with the polysaccharide biosynthesis domain. Like the predicted human secondary structure, YPL225W is experimentally verified to also contain four alpha helices and two beta sheets within the biosynthesis domain. Like CXorf26, YPL225W function in yeast is unknown, but based on co-purification experiments it may interact with ribosomes since many of its 18 interacting proteins were related to RNA and ribosomes. There were also multiple proteins involved with RNA polymerase, which is involved in the cellular process of transcription. Furthermore, multiple proteins were involved in ubiquitination. Some of the interacting yeast proteins with the higher interaction scores were UBI4, RPB8, SRO9, and NAB2.
Interacting proteins
Potential interacting proteins were identified using the tools provided at the I2D Interlogous Interaction Database and the STRING 9.0 program. Although more proteins were predicted, those shown below had the highest scores and showed the greatest possibility of relating to potential CXorf26 function.
SMAD2, PHB, and CTNNB1 were found in an experiment investigating transcriptional factor networks. The BABAM1 interaction was found in both databases using an anti-tag coimmunoprecipitation assay while POLR2H was based on a tandem affinity purification assay using the yeast homolog, YPL225W.
| Interacting Protein | Accession number | Protein Function |
|---|---|---|
| SMAD2 | AAC39657.1 | Part of family acting as signal transducer and transcriptional modulator |
| PHB | CAG46507.1 | Evolutionary conserved, ubiquitously expressed, negative regulator of cell proliferation |
| CTNNB1 | NP_001091679.1 | Catenin associated, part of protein complex that constructs adherens junctions |
| BABAM1 | NP_001028721.1 | Part of complex that recognizes Lys-63 ubiquinated histones |
| BRIX1 | NP_060791.3 | Required for biogenesis of 60s large eukaryotic ribosomal subunit |
| POLR2H | NP_006223.2 | Encodes essentential subunit of RNA Polymerase II |
References
References
- (2005). "Ubiquitin and control of transcription". Essays Biochem..
- (August 2010). "Emerging role of Lys-63 ubiquitination in protein kinase and phosphatase activation and cancer development". Oncogene.
- {{GeneCard. CXorf26
- [https://www.ncbi.nlm.nih.gov/IEB/Research/Acembly/av.cgi?db=human&term=cxorf26 Aceview Gene Annotation]
- Stevenson RE. (2000). "[[GeneReviews]]". University of Washington.
- {{Uniprot. Q96MV8
- (2008). "A comprehensive functional analysis of tissue specificity of human gene expression". BMC Biol..
- [https://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS3510:224177_s_at] NCBI GEO Profile GDS3510: Claudin-1 overexpression effect on lung adenocarcinoma cell line
- (January 2009). "Claudin-1 is a metastasis suppressor and correlates with clinical outcome in lung adenocarcinoma". Am. J. Respir. Crit. Care Med..
- [Ensembl Genome Browser http://useast.ensembl.org/Homo_sapiens/Gene/Summary?g=ENSG00000102390;r=X:75392771-75398039]
- "SoftBerry FGENESH".
- Genomatix: Eldorado Genome Annotation and Browser [www.genomatix.de]
- (1999). "PSORT: A program for detecting sorting signals in proteins and predicting their subcellular localization". Trends in Biochemical Sciences.
- (October 2003). "Global analysis of protein localization in budding yeast". Nature.
- [http://workbench.sdsc.edu/] SDSC BiologyWorkbench: TMAP
- [https://blast.ncbi.nlm.nih.gov/Blast.cgi NCBI BLAST Assembled RefSeq Genomes]
- [https://www.ncbi.nlm.nih.gov/Structure/cdd/cddsrv.cgi?ascbin=8&maxaln=10&seltype=2&uid=191055 NCBI Conserved Domain Database]
- (December 2000). "Heparin and heparan sulfate: biosynthesis, structure and function". Curr Opin Chem Biol.
- (November 2001). "Enzyme interactions in heparan sulfate biosynthesis: uronosyl 5-epimerase and 2-O-sulfotransferase interact in vivo". Proc. Natl. Acad. Sci. U.S.A..
- ExPASy Tools [http://expasy.org/tools/]
- [A Novel Solution NMR Structure of Protein yst0336 from Saccharomyces cerevisiae https://www.ncbi.nlm.nih.gov/Structure/mmdb/mmdbsrv.cgi?uid=61478&Dopt=s]
- [ExPASy Tools: Peptide Cutter http://expasy.org/tools/]
- [ExPASy Tools: NetGlycate http://expasy.org/tools/]
- [NCBI BLAST Alignment Tool https://blast.ncbi.nlm.nih.gov/Blast.cgi]
- [http://workbench.sdsc.edu/ SDSC Biology Workbench tools]
- Wu B, Yee A, Fares C, Lemak A, Gutmanas A, Semest A, Arrowsmith CH. [A Novel Solution NMR Structure of Protein yst0336 from Saccharomyces cerevisiae https://www.ncbi.nlm.nih.gov/Structure/mmdb/mmdbsrv.cgi?uid=61478&Dopt=s]
- [https://archive.today/20120712181745/http://ophid.utoronto.ca/ophidv2.201/ForwardingServlet?proteins=Q9BVG4&inputFormat=SWISSPROT_ID&ophidVersion=1.9&ophidOrganism=HUMAN&outputFormat=htmlOutput] I2D Protein Interaction Database
- [http://string-db.org/] STRING 9.0 Protein Interaction Predictor
- (February 2010). "A comprehensive resource of interacting protein regions for refining human transcription factor networks". PLOS ONE.
- (July 2009). "Defining the human deubiquitinating enzyme interaction landscape". Cell.
- (March 2006). "Global landscape of protein complexes in the yeast Saccharomyces cerevisiae". Nature.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about PBDC1 — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report