From Surf Wiki (app.surf) — the open knowledge base
Novobiocin
Chemical compound
Chemical compound
| Drugs.com = | elimination_half-life = 6 hours
Novobiocin, also known as albamycin, is an aminocoumarin antibiotic that is produced by the actinomycete Streptomyces niveus, which is a heterotypic synonym for S. spheroides a member of the class Actinomycetia. Other aminocoumarin antibiotics include clorobiocin and coumermycin A1. Novobiocin was first reported in the mid-1950s (then called streptonivicin).
Clinical use
It is active against Staphylococcus epidermidis and may be used to differentiate S. epidermidis from coagulase-negative Staphylococcus saprophyticus, which is resistant to novobiocin, in culture.
Novobiocin was licensed for clinical use under the tradename Albamycin (Upjohn) in the 1960s. Its efficacy has been demonstrated in preclinical and clinical trials. The oral form of the drug has since been withdrawn from the market due to lack of efficacy. A combination product of novobiocin and tetracycline, sold by Upjohn under brand names such as Panalba and Albamycin-T, was in particular the subject of intense FDA scrutiny before it was finally taken off the market. Novobiocin is an effective antistaphylococcal agent used in the treatment of MRSA.
Mechanism of action
The molecular basis of action of novobiocin, and other related drugs clorobiocin and coumermycin A1, has been examined. Aminocoumarins are very potent inhibitors of bacterial DNA gyrase and work by targeting the GyrB subunit of the enzyme involved in energy transduction. Novobiocin as well as the other aminocoumarin antibiotics act as competitive inhibitors of the ATPase reaction catalysed by GyrB. The potency of novobiocin is considerably higher than that of the fluoroquinolones that also target DNA gyrase, but at a different site on the enzyme. The GyrA subunit is involved in the DNA nicking and ligation activity.
Novobiocin has been shown to weakly inhibit the C-terminus of the eukaryotic Hsp90 protein (high micromolar IC50). Modification of the novobiocin scaffold has led to more selective Hsp90 inhibitors. Novobiocin has also been shown to bind and activate the Gram-negative lipopolysaccharide transporter LptBFGC.
The ATP binding pocket of polymerase theta is blocked by novobiocin resulting in a loss of ATPase activity. This results in the loss of microhomology-mediated end joining as a pathway for homologous recombination deficient cells to circumvent DNA damaging agents. The action of novobiocin is syngeristic with PARP inhibitors for reducing tumor size in a mouse model.
Structure
Novobiocin is an aminocoumarin. Novobiocin may be divided up into three entities; a benzoic acid derivative, a coumarin residue, and the sugar novobiose. The overlap of the coumarin and ATP-binding sites is consistent with aminocoumarins being competitive inhibitors of the ATPase activity.
Structure–activity relationship
In structure activity relationship experiments it was found that removal of the carbamoyl group located on the novobiose sugar lead to a dramatic decrease in inhibitory activity of novobiocin.
Biosynthesis
This aminocoumarin antibiotic consists of three major substituents. The 3-dimethylallyl-4-hydroxybenzoic acid moiety, known as ring A, is derived from prephenate and dimethylallyl pyrophosphate. The aminocoumarin moiety, known as ring B, is derived from L-tyrosine. The final component of novobiocin is the sugar derivative L-noviose, known as ring C, which is derived from glucose-1-phosphate. The biosynthetic gene cluster for novobiocin was identified by Heide and coworkers in 1999 (published 2000) from Streptomyces spheroides NCIB 11891. They identified 23 putative open reading frames (ORFs) and more than 11 other ORFs that may play a role in novobiocin biosynthesis.
The biosynthesis of ring A (see Fig. 1) begins with prephenate which is a derived from the shikimic acid biosynthetic pathway. The enzyme NovF catalyzes the decarboxylation of prephenate while simultaneously reducing nicotinamide adenine dinucleotide phosphate (NADP+) to produce NADPH. Following this NovQ catalyzes the electrophilic substitution of the phenyl ring with dimethylallyl pyrophosphate (DMAPP) otherwise known as prenylation. DMAPP can come from either the mevalonic acid pathway or the deoxyxylulose biosynthetic pathway. Next the 3-dimethylallyl-4-hydroxybenzoate molecule is subjected to two oxidative decarboxylations by NovR and molecular oxygen. NovR is a non-heme iron oxygenase with a unique bifunctional catalysis. In the first stage both oxygens are incorporated from the molecular oxygen while in the second step only one is incorporated as determined by isotope labeling studies. This completes the formation of ring A.
The biosynthesis of ring B (see Fig. 2) begins with the natural amino acid L-tyrosine. This is then adenylated and thioesterified onto the peptidyl carrier protein (PCP) of NovH by ATP and NovH itself. NovI then further modifies this PCP bound molecule by oxidizing the β-position using NADPH and molecular oxygen. NovJ and NovK form a heterodimer of J2K2 which is the active form of this benzylic oxygenase. This process uses NADP+ as a hydride acceptor in the oxidation of the β-alcohol. This ketone will prefer to exist in its enol tautomer in solution. Next a still unidentified protein catalyzes the selective oxidation of the benzene (as shown in Fig. 2). Upon oxidation this intermediate will spontaneously lactonize to form the aromatic ring B and lose NovH in the process.

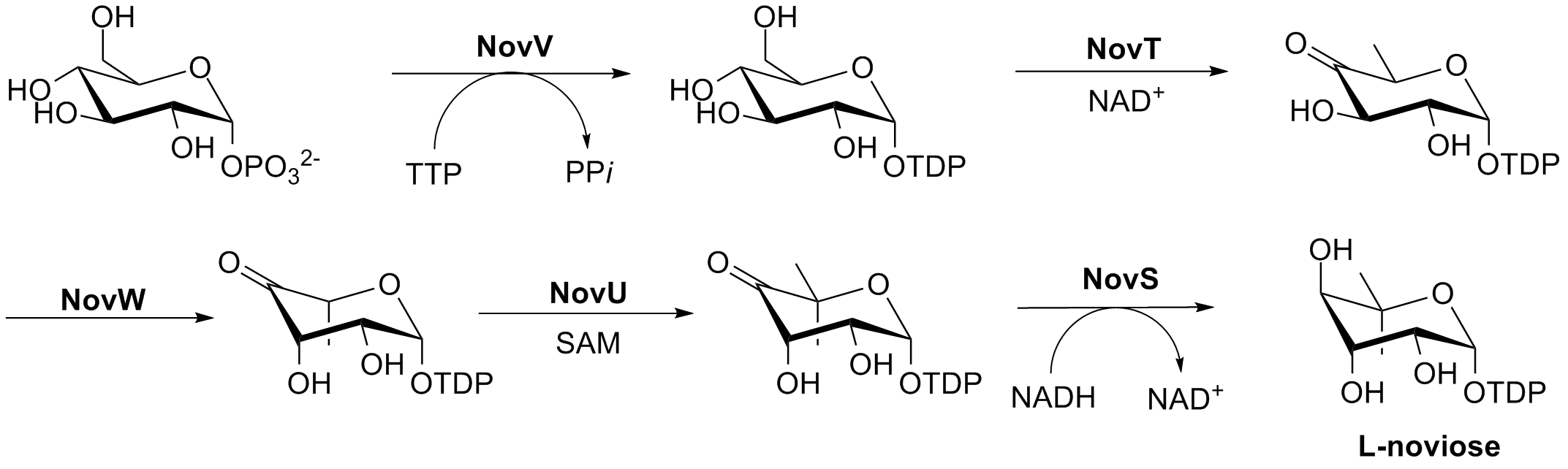
The biosynthesis of L-noviose (ring C) is shown in Fig. 3. This process starts from glucose-1-phosphate where NovV takes dTTP and replaces the phosphate group with a dTDP group. NovT then oxidizes the 4-hydroxy group using NAD+. NovT also accomplishes a dehydroxylation of the 6 position of the sugar. NovW then epimerizes the 3 position of the sugar. The methylation of the 5 position is accomplished by NovU and S-adenosyl methionine (SAM). Finally NovS reduces the 4 position again to achieve epimerization of that position from the starting glucose-1-phosphate using NADH.
Rings A, B, and C are coupled together and modified to give the finished novobiocin molecule. Rings A and B are coupled together by the enzyme NovL using ATP to diphosphorylate the carboxylate group of ring A so that the carbonyl can be attacked by the amine group on ring B. The resulting compound is methylated by NovO and SAM prior to glycosylation. NovM adds ring C (L-noviose) to the hydroxyl group derived from tyrosine with the loss of dTDP. Another methylation is accomplished by NovP and SAM at the 4 position of the L-noviose sugar. This methylation allows NovN to carbamylate the 3 position of the sugar as shown in Fig. 4 completing the biosynthesis of novobiocin.
]]
References
References
- Komaki, Hisayuki. (Mar 2023). "Recent progress of reclassification of the genus Streptomyces". Microorganisms.
- (December 1955). "Structural studies on streptonivicin, a new antibiotic". Journal of the American Chemical Society.
- (February 1956). "Streptonivicin, a new antibiotic. I. Discovery and biologic studies". Antibiotics & Chemotherapy.
- (1983). "Effect of Different Media on Determination of Novobiocin Resistance for Differentiation of Coagulase-Negative Staphylococci". Journal of Clinical Microbiology.
- (2007). "Intrinsic Novobiocin Resistance in ''Staphylococcus saprophyticus''". Antimicrobial Agents and Chemotherapy.
- (November 1995). "Antibiotics and prevention of microbial colonization of catheters". Antimicrobial Agents and Chemotherapy.
- (January 1998). "A prospective crossover randomized trial of novobiocin and rifampin prophylaxis for the prevention of intravascular catheter infections in cancer patients treated with interleukin-2". Cancer.
- (19 January 2011). "Determination That ALBAMYCIN (Novobiocin Sodium) Capsule, 250 Milligrams, Was Withdrawn From Sale for Reasons of Safety or Effectiveness". The Federal Register.
- (August 1969). "FDA and Panalba: A Conflict of Commercial, Therapeutic Goals?". Science.
- "Upjohn's Shuck and Jive Routine". Mother Jones and the Foundation for National Progress.
- (June 1993). "Randomized double-blinded trial of rifampin with either novobiocin or trimethoprim-sulfamethoxazole against methicillin-resistant Staphylococcus aureus colonization: prevention of antimicrobial resistance and effect of host factors on outcome". Antimicrobial Agents and Chemotherapy.
- (August 1993). "The interaction between coumarin drugs and DNA gyrase". Molecular Microbiology.
- (February 1999). "DNA gyrase as a drug target". Biochemical Society Transactions.
- (August 1996). "Molecular mechanisms of drug inhibition of DNA gyrase". BioEssays.
- (2003). "The ATP-binding site of type II topoisomerases as a target for antibacterial drugs". Current Topics in Medicinal Chemistry.
- (September 2005). "Hsp90 inhibitors identified from a library of novobiocin analogues". Journal of the American Chemical Society.
- (June 2018). "Novobiocin Enhances Polymyxin Activity by Stimulating Lipopolysaccharide Transport". Journal of the American Chemical Society.
- (December 2017). "The Antibiotic Novobiocin Binds and Activates the ATPase That Powers Lipopolysaccharide Transport". Journal of the American Chemical Society.
- (June 2021). "A first-in-class polymerase theta inhibitor selectively targets homologous-recombination-deficient tumors". Nature Cancer.
- (May 1997). "The high-resolution crystal structure of a 24-kDa gyrase B fragment from E. coli complexed with one of the most potent coumarin inhibitors, clorobiocin". Proteins.
- (April 2006). "Structure-activity relationships of aminocoumarin-type gyrase and topoisomerase IV inhibitors obtained by combinatorial biosynthesis". Antimicrobial Agents and Chemotherapy.
- (May 2000). "Identification of the novobiocin biosynthetic gene cluster of Streptomyces spheroides NCIB 11891". Antimicrobial Agents and Chemotherapy.
- (March 2003). "CloQ, a prenyltransferase involved in clorobiocin biosynthesis". Proceedings of the National Academy of Sciences of the United States of America.
- (August 2003). "CloR, a bifunctional non-heme iron oxygenase involved in clorobiocin biosynthesis". The Journal of Biological Chemistry.
- (April 2001). "Coumarin formation in novobiocin biosynthesis: beta-hydroxylation of the aminoacyl enzyme tyrosyl-S-NovH by a cytochrome P450 NovI". Chemistry & Biology.
- (September 2005). "NovJ/NovK catalyze benzylic oxidation of a beta-hydroxyl tyrosyl-S-pantetheinyl enzyme during aminocoumarin ring formation in novobiocin biosynthesis". Biochemistry.
- (April 2005). "Functional characterizations of novWUS involved in novobiocin biosynthesis from Streptomyces spheroides". Archives of Biochemistry and Biophysics.
- (November 2005). "CouO and NovO: C-methyltransferases for tailoring the aminocoumarin scaffold in coumermycin and novobiocin antibiotic biosynthesis". Biochemistry.
- (January 2004). "Characterization of NovP and NovN: completion of novobiocin biosynthesis by sequential tailoring of the noviosyl ring". Angewandte Chemie.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Novobiocin — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report