From Surf Wiki (app.surf) — the open knowledge base
Microbiota
Community of microorganisms
Community of microorganisms

Microbiota are the range of microorganisms that may be commensal, mutualistic, or pathogenic found in and on all multicellular organisms, including plants. Microbiota include bacteria, archaea, protists, fungi, and viruses, and have been found to be crucial for immunologic, hormonal, and metabolic homeostasis of their host.
The term microbiome describes either the collective genomes of the microbes that reside in an ecological niche or else the microbes themselves.
The microbiome and host emerged during evolution as a synergistic unit from epigenetics and genetic characteristics, sometimes collectively referred to as a holobiont. The presence of microbiota in human and other metazoan guts has been critical for understanding the co-evolution between metazoans and bacteria. Microbiota play key roles in the intestinal immune and metabolic responses via their fermentation product (short-chain fatty acid), acetate.
Introduction

All plants and animals, from simple life forms to humans, live in close association with microbial organisms. Several advances have driven the perception of microbiomes, including:
- the ability to perform genomic and gene expression analyses of single cells and of entire microbial communities in the disciplines of metagenomics and metatranscriptomics
- databases accessible to researchers across multiple disciplines
- methods of mathematical analysis suitable for complex data sets
Biologists discovered that microbes make up an important part of an organism's phenotype, far beyond the occasional symbiotic case study.
Types of microbe-host relationships
Commensalism, a concept developed by Pierre-Joseph van Beneden (1809–1894), a Belgian professor at the University of Louvain during the nineteenth century is central to the microbiome, where microbiota colonize a host in a non-harmful coexistence. The relationship with their host is called mutualistic when organisms perform tasks that are known to be useful for the host, parasitic, when disadvantageous to the host. Other authors define a situation as mutualistic where both benefit, and commensal, where the unaffected host benefits the symbiont.
Acquisition and change
The initial acquisition of microbiota in animals from mammalians to marine sponges is at birth, and may even occur through the germ cell line. In plants, the colonizing process can be initiated below ground in the root zone, around the germinating seed, the spermosphere, or originate from the above ground parts, the phyllosphere and the flower zone or anthosphere. Clinically, new microbiota can be acquired through fecal microbiota transplant to treat infections such as chronic C. difficile infection.
Microbiota by host
Humans
Main article: Human microbiota
The human microbiota includes bacteria, fungi, archaea and viruses. Micro-animals which live on the human body are excluded. The human microbiome refers to their collective genomes.
Humans are colonized by many microorganisms; the traditional estimate was that humans live with ten times more non-human cells than human cells; more recent estimates have lowered this to 3:1 and even to about 1:1 by number (1:350 by mass).
In fact, these are so small that there are around 100 trillion microbiota on the human body, around 39 trillion by revised estimates, with only 0.2 kg of total mass in a "reference" 70 kg human body.
The Human Microbiome Project sequenced the genome of the human microbiota, focusing particularly on the microbiota that normally inhabit the skin, mouth, nose, digestive tract, and vagina. It reached a milestone in 2012 when it published initial results.
Non-human animals
- Amphibians have microbiota on their skin. Some species are able to carry a fungus named Batrachochytrium dendrobatidis, which in others can cause a deadly infection Chytridiomycosis depending on their microbiome, resisting pathogen colonization or inhibiting their growth with antimicrobial skin peptides.
- Newborn marsupials are born with histologically immature immune tissues and unable to mount their own specific immune defence. They are therefore heavily reliant on their mother's immune system and the milk for their protection. Most marsupials have pouches, and their own microbiota changes throughout the reproductive stages: oestrus, birth/oestrus, and post-oestrus. Some pouch and skin secretions have had antimicrobial peptides identified, that presumably support the young at this vulnerable time.
- In mammals, herbivores such as cattle depend on their rumen microbiome to convert cellulose into proteins, short chain fatty acids, and gases. Culture methods cannot provide information on all microorganisms present. Comparative metagenomic studies yielded the surprising result that individual cattle possess markedly different community structures, predicted phenotype, and metabolic potentials,{{cite journal |doi-access=free }} even though they were fed identical diets, were housed together, and were apparently functionally identical in their utilization of plant cell wall resources.
- Mice have become the most studied mammalian regarding their microbiomes. The gut microbiota have been studied in relation to allergic airway disease, obesity, gastrointestinal diseases and diabetes. Perinatal shifting of microbiota through low dose antibiotics can have long-lasting effects on future susceptibility to allergic airway disease. The frequency of certain subsets of microbes has been linked to disease severity. The presence of specific microbes early in postnatal life, instruct future immune responses. In gnotobiotic mice certain gut bacteria were found to transmit a particular phenotype to recipient germ-free mice, that promoted accumulation of colonic regulatory T cells, and strains that modulated mouse adiposity and cecal metabolite concentrations. This combinatorial approach enables a systems-level understanding of microbial contributions to human biology. But also other mucoide tissues as lung and vagina have been studied in relation to diseases such as asthma, allergy and vaginosis.
- Insects have their own microbiomes. For example, leaf-cutter ants form huge underground colonies harvesting hundreds of kilograms of leaves each year and are unable to digest the cellulose in the leaves directly. They maintain fungus gardens as the colony's primary food source. While the fungus itself does not digest cellulose, a microbial community containing a diversity of bacteria is doing so. Analysis of the microbial population's genome revealed many genes with a role in cellulose digestion. This microbiome's predicted carbohydrate-degrading enzyme profile is similar to that of the bovine rumen, but the species composition is almost entirely different. Gut microbiota of the fruit fly can affect the way its gut looks, by impacting epithelial renewal rate, cellular spacing, and the composition of different cell types in the epithelium. When the moth Spodoptera exigua is infected with baculovirus immune-related genes are downregulated and the amount of its gut microbiota increases. In the dipteran intestine, enteroendocrine cells sense the gut microbiota-derived metabolites and coordinate antibacterial, mechanical, and metabolic branches of the host intestinal innate immune response to the commensal microbiota.
- Fish have their own microbiomes, including the short-lived species Nothobranchius furzeri (turquoise killifish). Transferring the gut microbiota from young killfish into middle-aged killifish significantly extends the lifespans of the middle-aged killfish.
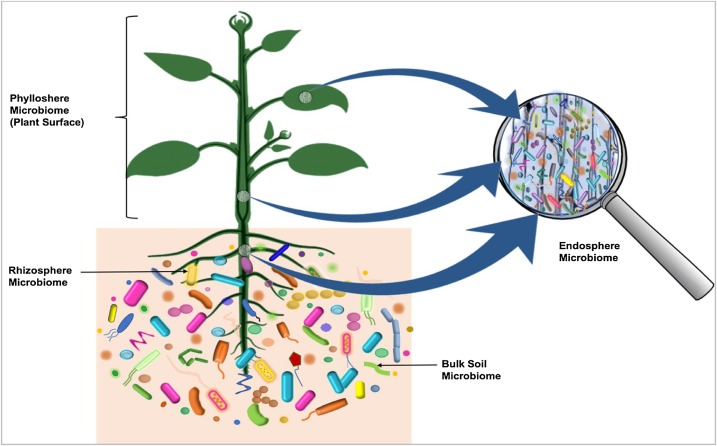
Plants

The plant microbiome was recently discovered to originate from the seed. Microorganism which are transmitted via seed migrate into the developing seedling in a specific route in which certain community move to the leaves and others to the roots. In the diagram on the right, microbiota colonizing the rhizosphere, entering the roots and colonizing the next tuber generation via the stolons, are visualized with a red color. Bacteria present in the mother tuber, passing through the stolons and migrating into the plant as well as into the next generation of tubers are shown in blue.
- The soil is the main reservoir for bacteria that colonize potato tubers
- Bacteria are recruited from the soil more or less independent of the potato variety
- Bacteria might colonize the tubers predominantly from the inside of plants via the stolon
- The bacterial microbiota of potato tubers consists of bacteria transmitted from one tuber generation to the next and bacteria recruited from the soil colonize potato plants via the root.

Plants are attractive hosts for microorganisms since they provide a variety of nutrients. Microorganisms on plants can be epiphytes (found on the plants) or endophytes (found inside plant tissue). Oomycetes and fungi have, through convergent evolution, developed similar morphology and occupy similar ecological niches. They develop hyphae, threadlike structures that penetrate the host cell. In mutualistic situations the plant often exchanges hexose sugars for inorganic phosphate from the fungal symbiont. It is speculated that such very ancient associations have aided plants when they first colonized land.{{cite journal | doi-access=free | doi-access=free
Research
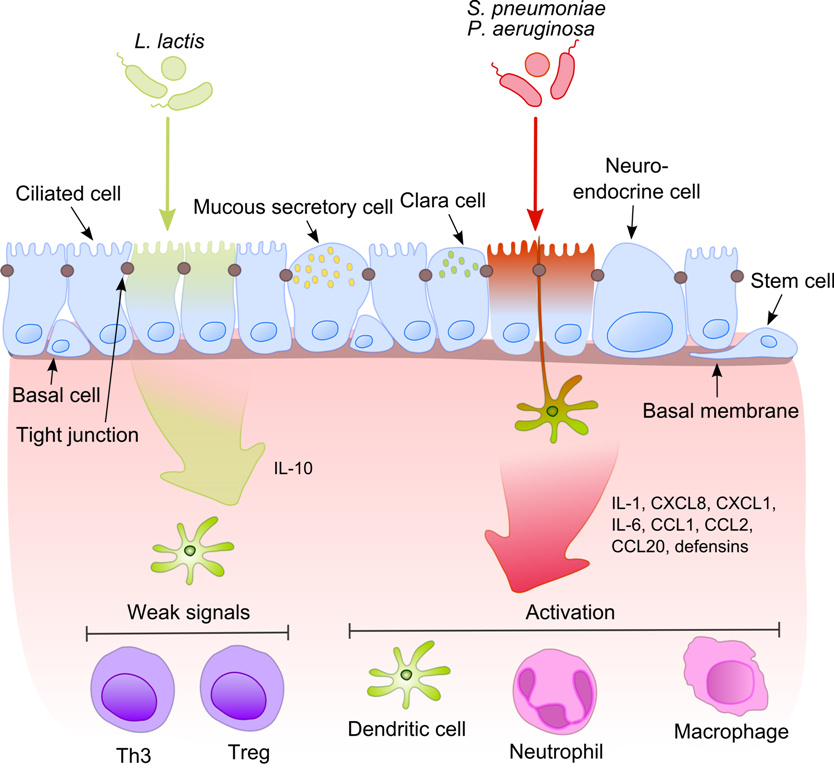
The symbiotic relationship between a host and its microbiota is under laboratory research for how it may shape the immune system of mammals. In many animals, the immune system and microbiota may engage in "cross-talk" by exchanging chemical signals, which may enable the microbiota to influence immune reactivity and targeting. Bacteria can be transferred from mother to child through direct contact and after birth. As the infant microbiome is established, commensal bacteria quickly populate the gut, prompting a range of immune responses and "programming" the immune system with long-lasting effects. The bacteria are able to stimulate lymphoid tissue associated with the gut mucosa, which enables the tissue to produce antibodies for pathogens that may enter the gut.
The human microbiome may play a role in the activation of toll-like receptors in the intestines, a type of pattern recognition receptor host cells use to recognize dangers and repair damage. Pathogens can influence this coexistence leading to immune dysregulation including and susceptibility to diseases, mechanisms of inflammation, immune tolerance, and autoimmune diseases.
Co-evolution of microbiota
Main article: Hologenome theory of evolution

Organisms evolve within ecosystems so that the change of one organism affects the change of others. The hologenome theory of evolution proposes that an object of natural selection is not the individual organism, but the organism together with its associated organisms, including its microbial communities.
Coral reefs. The hologenome theory originated in studies on coral reefs. Coral reefs are the largest structures created by living organisms, and contain abundant and highly complex microbial communities. Over the past several decades, major declines in coral populations have occurred. Climate change, water pollution and over-fishing are three stress factors that have been described as leading to disease susceptibility. Over twenty different coral diseases have been described, but of these, only a handful have had their causative agents isolated and characterized. Coral bleaching is the most serious of these diseases. In the Mediterranean Sea, the bleaching of Oculina patagonica was first described in 1994 and shortly determined to be due to infection by Vibrio shiloi. From 1994 to 2002, bacterial bleaching of O. patagonica occurred every summer in the eastern Mediterranean. Surprisingly, however, after 2003, O. patagonica in the eastern Mediterranean has been resistant to V. shiloi infection, although other diseases still cause bleaching. The surprise stems from the knowledge that corals are long lived, with lifespans on the order of decades, and do not have adaptive immune systems. Their innate immune systems do not produce antibodies, and they should seemingly not be able to respond to new challenges except over evolutionary time scales.
The puzzle of how corals managed to acquire resistance to a specific pathogen led to a 2007 proposal, that a dynamic relationship exists between corals and their symbiotic microbial communities. It is thought that by altering its composition, the holobiont can adapt to changing environmental conditions far more rapidly than by genetic mutation and selection alone. Extrapolating this hypothesis to other organisms, including higher plants and animals, led to the proposal of the hologenome theory of evolution.
the hologenome theory was still being debated.{{cite journal | doi-access=free}} A major criticism has been the claim that V. shiloi was misidentified as the causative agent of coral bleaching, and that its presence in bleached O. patagonica was simply that of opportunistic colonization.{{cite journal | doi-access=free
Research methods
Targeted amplicon sequencing
Targeted amplicon sequencing relies on having some expectations about the composition of the community that is being studied. In target amplicon sequencing a phylogenetically informative marker is targeted for sequencing. Such a marker should be present in ideally all the expected organisms. It should also evolve in such a way that it is conserved enough that primers can target genes from a wide range of organisms while evolving quickly enough to allow for finer resolution at the taxonomic level. A common marker for human microbiome studies is the gene for bacterial 16S rRNA (i.e. "16S rDNA", the sequence of DNA which encodes the ribosomal RNA molecule).{{Cite journal
Targeted studies of eukaryotic and viral communities are limited{{Cite book | doi-access = free
After the amplicons are sequenced, molecular phylogenetic methods are used to infer the composition of the microbial community. This can be done through clustering methodologies, by clustering the amplicons into operational taxonomic units (OTUs); or alternatively with denoising methodologies, identifying amplicon sequence variants (ASVs).
Phylogenetic relationships are then inferred between the sequences. Due to the complexity of the data, distance measures such as UniFrac distances are usually defined between microbiome samples, and downstream multivariate methods are carried out on the distance matrices. An important point is that the scale of data is extensive, and further approaches must be taken to identify patterns from the available information. Tools used to analyze the data include VAMPS, QIIME,{{Cite journal
Metagenomic sequencing
Main article: Metagenomics
Metagenomics is also used extensively for studying microbial communities.{{Cite journal
Despite the fact that metagenomics is limited by the availability of reference sequences, one significant advantage of metagenomics over targeted amplicon sequencing is that metagenomics data can elucidate the functional potential of the community DNA.{{Cite journal | article-number = 386 | doi-access = free
RNA and protein-based approaches
Metatranscriptomics studies have been performed to study the gene expression of microbial communities through methods such as the pyrosequencing of extracted RNA.{{Cite journal| last1 = Shi | first1 = Y.| last2 = Tyson | first2 = G. W.
Projects
The Human Microbiome Project launched in 2008 was a United States National Institutes of Health initiative to identify and characterize microorganisms found in both healthy and diseased humans. The five-year project, best characterized as a feasibility study with a budget of $115 million, tested how changes in the human microbiome are associated with human health or disease.
The Earth Microbiome Project (EMP) is an initiative to collect natural samples and analyze the microbial community around the globe. Microbes are highly abundant, diverse and have an important role in the ecological system. Yet , it was estimated that the total global environmental DNA sequencing effort had produced less than 1 percent of the total DNA found in a liter of seawater or a gram of soil, and the specific interactions between microbes are largely unknown. The EMP aims to process as many as 200,000 samples in different biomes, generating a complete database of microbes on earth to characterize environments and ecosystems by microbial composition and interaction. Using these data, new ecological and evolutionary theories can be proposed and tested.
Privacy issues
Microbial DNA inhabiting a person's human body can uniquely identify the person. A person's privacy may be compromised if the person anonymously donated microbe DNA data. Their medical condition and identity could be revealed.
References
References
- (2020). "Plant microbiome–an account of the factors that shape community composition and diversity". Current Plant Biology.
- (2019). "The battle within: interactions of bacteriophages and bacteria in the gastrointestinal tract". [[Cell Host & Microbe]].
- (2009). "The NIH Human Microbiome Project". NIH HMP Working Group.
- (2005). "Host-Bacterial Mutualism in the Human Intestine". [[Science (journal).
- (2007). "The Human Microbiome Project". [[Nature (journal).
- (2006). "Ecological and Evolutionary Forces Shaping Microbial Diversity in the Human Intestine". [[Cell (journal).
- (2016). "Microbiome, holobiont and the net of life". [[Critical Reviews in Microbiology]].
- (2013). "Symbiogenesis: The holobiont as a unit of evolution". [[International Microbiology]].
- (2017). "The human microbiome in evolution". BMC Biology.
- (2014). "Rapid changes in the gut microbiome during human evolution". Proceedings of the National Academy of Sciences.
- (2021). "Microbiota-derived acetate activates intestinal innate immunity via the Tip60 histone acetyltransferase complex". [[Immunity (journal).
- (2015). "Cross-kingdom similarities in microbiome functions". The ISME Journal.
- (2011). "Metaorganisms as the new frontier". Zoology.
- Poreau B., ''[http://www.theses.fr/2014LYO10114 Biologie et complexité : histoire et modèles du commensalisme]''. PhD Dissertation, University of Lyon, France, 2014.
- (Sep 2013). "Gut bacteria in health and disease". Gastroenterol Hepatol (N Y).
- Copeland, CS. (Sep–Oct 2017). "The World Within Us". Healthcare Journal of New Orleans.
- (Nov 2015). "Stability and succession of the rhizosphere microbiota depends upon plant type and soil composition". ISME J..
- Copeland, CS. (19 April 2019). "What is Clostridium difficile?".
- (2013). ["Prescott's Microbiology"]({{google books). McGraw Hill.
- American Academy of Microbiology [http://academy.asm.org/index.php/faq-series/5122-humanmicrobiome FAQ: Human Microbiome] {{Webarchive. link. (2016-12-31 January 2014)
- (2014). "Ten Times More Microbial Cells than Body Cells in Humans?". Microbe Magazine.
- (2016). "Scientists bust myth that our bodies have more bacteria than human cells". Nature.
- (Jan 2016). "Are We Really Vastly Outnumbered? Revisiting the Ratio of Bacterial to Host Cells in Humans". Cell.
- (2016-08-19). "Revised Estimates for the Number of Human and Bacteria Cells in the Body". PLOS Biology.
- "On and in You." Micropia, https://www.micropia.nl/en/discover/stories/on-and-in-you/#:~:text=They're%20on%20you%2C%20in,re%20known%20as%20human%20microbiota.
- (13 June 2012). "NIH Human Microbiome Project defines normal bacterial makeup of the body". NIH News.
- (Jan 2016). "Microbiome Variation Across Amphibian Skin Regions: Implications for Chytridiomycosis Mitigation Efforts". Microb. Ecol..
- (2007). "Innate immune defenses of amphibian skin: antimicrobial peptides and more". Animal Conservation.
- (2000). "Development of the immune system and immunological protection in marsupial pouch young". Developmental and Comparative Immunology.
- (2020). "Marsupial and monotreme milk – a review of its nutrients and immune properties". PeerJ.
- (1998). "The effect of oestrus and the presence of pouch young on aerobic bacteria isolated from the pouch of the tammar wallaby, ''Macropus eugenii''". Comparative Immunology Microbiology and Infectious Diseases.
- Russell SL, Gold MJ. (May 2012). "Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma". EMBO Rep..
- (Aug 2014). "Perinatal antibiotic-induced shifts in gut microbiota have differential effects on inflammatory lung diseases". J Allergy Clin Immunol.
- (Dec 2006). "An obesity-associated gut microbiome with increased capacity for energy harvest". Nature.
- (Jan 2014). "Identifying gut microbe-host phenotype relationships using combinatorial communities in gnotobiotic mice.". Sci. Transl. Med..
- (2013). "The murine lung microbiome in relation to the intestinal and vaginal bacterial communities". BMC Microbiol.
- (2010). "An Insect Herbivore Microbiome with High Plant Biomass-Degrading Capacity". PLOS Genet.
- (2014). "Microbiota-Induced Changes in Drosophila melanogaster Host Gene Expression and Gut Morphology". mBio.
- (May 2013). "Increase in Gut Microbiota after Immune Suppression in Baculovirus-infected Larvae". PLOS Pathog.
- (2020-02-01). "Microbial Control of Intestinal Homeostasis via Enteroendocrine Cell Innate Immune Signaling". Trends in Microbiology.
- (2019). "The influence of the microbiota on immune development, chronic inflammation, and cancer in the context of aging". Microbial Cell.
- Buchholz, F., Antonielli, L., Kostić, T., Sessitsch, A. and Mitter, B. (2019) "The bacterial community in potato is recruited from soil and partly inherited across generations". ''PLOS One'', '''14'''(11): e0223691. {{doi. 10.1371/journal.pone.0223691. 50px]] Material was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ Creative Commons Attribution 4.0 International License].
- (2021). "Experimental evidence of microbial inheritance in plants and transmission routes from seed to phyllosphere and root". Environmental Microbiology.
- Berlec, Aleš. (2012-09-01). "Novel techniques and findings in the study of plant microbiota: Search for plant probiotics". Plant Science.
- (2008-12-01). "Phyllosphere microbiology with special reference to diversity and plant genotype". Journal of Applied Microbiology.
- (2001). "Molecular basis of plant growth promotion and biocontrol by rhizobacteria". Current Opinion in Plant Biology.
- (30 June 2015). "Immune–microbiota interactions in health and disease". Clinical Immunology.
- (2010). "Coordination of tolerogenic immune responses by the commensal microbiota". Journal of Autoimmunity.
- (2012). "Microbial-immune cross-talk and regulation of the immune system". Immunology.
- (2016). "Microbes drive evolution of animals and plants: the hologenome concept". mBio.
- (19 July 2017). "Regulation of inflammation by microbiota interactions with the host". Nature Immunology.
- (2014). "Reciprocity in microbiome and immune system interactions and its implications in disease and health". Inflamm Allergy Drug Targets.
- (2009). "Coral bleaching: the role of the host". Trends in Ecology and Evolution.
- (2007). "The role of microorganisms in coral health, disease and evolution". Nature Reviews Microbiology.
- (May 2007). "A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria". Journal of Microbiological Methods.
- (December 2020). "Influence of 16S rRNA target region on the outcome of microbiome studies in soil and saliva samples". Scientific Reports.
- "VAMPS: The Visualization and Analysis of Microbial Population Structures". Bay Paul Center, MBL, Woods Hole.
- (July 2016). "DADA2: High-resolution sample inference from Illumina amplicon data". Nature Methods.
- Edgar, Robert C.. (2016-10-15). "UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing".
- (2005). "Comparative Metagenomics of Microbial Communities". Science.
- (2010). "A Primer on Metagenomics". PLOS Computational Biology.
- (2018-02-28). "Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen". Nature Communications.
- (2016). "NIH Human Microbiome Project". US National Institutes of Health, Department of Health and Human Services, US Government.
- (2010). "Meeting Report: The Terabase Metagenomics Workshop and the Vision of an Earth Microbiome Project". Standards in Genomic Sciences.
- (2011). "The importance of metagenomic surveys to microbial ecology: Or why Darwin would have been a metagenomic scientist". Microbial Informatics and Experimentation.
- magazine, Ewen. "Microbial DNA in Human Body Can Be Used to Identify Individuals".
- (2015). "Microbiomes raise privacy concerns". Nature.
- Yong, Ed. (2015-05-11). "Can The Microbes You Leave Behind Be Used to Identify You?". National Geographic.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Microbiota — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report