From Surf Wiki (app.surf) — the open knowledge base
Mediator (coactivator)
Multiprotein complex involved in transcription in eukaryotes

Multiprotein complex involved in transcription in eukaryotes

Mediator is a multiprotein complex that functions as a transcriptional coactivator in all eukaryotes. It was discovered in 1990 in the lab of Roger D. Kornberg, recipient of the 2006 Nobel Prize in Chemistry. Mediator interacts with transcription factors and RNA polymerase II. It mainly functions to transmit signals from the transcription factors to the polymerase.
Mediator complexes are variable at the evolutionary, compositional and conformational levels. Figure 1 shows only one "snapshot" of what a particular complex might comprise, but it is an inaccurate depiction of the conformation in vivo. During evolution, Mediator has complexified. The yeast Saccharomyces cerevisiae (a simple eukaryote) is thought to have up to 21 subunits in the core Mediator (exclusive of the CDK module), while mammals have up to 26.
Individual subunits can be absent or replaced by other subunits under different conditions. Also, there are many intrinsically disordered regions in Mediator proteins, which may contribute to the conformational flexibility seen both with and without other bound proteins or protein complexes. A more realistic model of Mediator without the CDK module is shown in Figure 2.
Mediator is required for successful transcription of genes by RNA polymerase II, and contacts the polymerase in the transcription preinitiation complex. A recent model showing the polymerase associating with Mediator without DNA is shown in Figure 3. Note that the different morphologies of Mediator do not necessarily mean that a particular model is correct; rather those differences may reflect the flexibility of Mediator as it interacts with other molecules. For example, after binding the enhancer and core promoter, the Mediator complex compositionally changes, dissociating the kinase module and associating with RNA polymerase II for transcriptional activation.
Mediator is located within the cell nucleus. It is required for successfully transcribing nearly all class II gene promoters in yeast. It works similarly in mammals. Mediator functions as a coactivator and binds to the C-terminal domain of RNA polymerase II holoenzyme, bridging this enzyme and transcription factors.
Structure
The yeast Mediator complex is approximately as massive as a small subunit of a eukaryotic ribosome. The yeast Mediator has 25 subunits, while the mammalian Mediator is slightly larger.
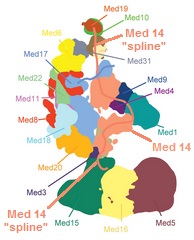
Mediator subunits have many intrinsically disordered regions called "splines", which may be important to allow the structural changes of Mediator that change the function of the complex. Figure 5 shows the splines of the MED14 subunit connecting a large portion of the complex together while still allowing flexibility.
Mediator complexes lacking a subunit have been found or produced. These smaller complexes can still function normally in some activity, but lack other capabilities. This indicates a somewhat independent function of some of the subunits while composing the larger complex.
Another example of structural variability is seen in vertebrates, in which 3 paralogues of subunits of the cyclin-dependent kinase (CDK) module have evolved by 3 independent gene duplication events followed by sequence divergence. These stable associations regulate gene expression in vivo, and are prevented by mutations in MED12 that produce the human disease FG syndrome. Thus, the structure of a Mediator complex can be augmented by RNA as well as proteinaceous transcription factors.
Function

Mediator was originally discovered because it was important for RNA polymerase II function, but it has many more functions than just interactions at the transcription start site.
RNA polymerase II–Mediator core initiation complex

Mediator is a crucial component for transcription initiation. Mediator interacts with the pre-initiation complex, composed of RNA Polymerase II and general transcription factors TFIIB, TFIID, TFIIE, TFIIF, and TFIIH to stabilize and initiate transcription. Studies of Mediator–RNA Pol II contacts in budding yeast showed the importance of TFIIB-Mediator contacts in the formation of the complex. Interactions of Mediator with TFIID in the initiation complex has been shown.
The structure of a core Mediator (cMed) while associated with a core pre-initiation complex was elucidated.
RNA synthesis
The preinitiation complex, which contains Mediator, transcription factors, a nucleosome and RNA polymerase II, is important for positioning the polymerase for the start of transcription. Before RNA synthesis starts, the polymerase dissociates from Mediator. This is seemingly via phosphorylation of the polymerase by a kinase. Importantly, Mediator and transcription factors do not dissociate from the DNA when the polymerase begins transcription. Rather, the complex remains at the promoter to recruit another RNA polymerase to begin another round of transcription.
There is some evidence to suggest that Mediator in Schizosaccharomyces pombe helps regulate RNA polymerase III (Pol III) transcripts of tRNAs. An independent report confirmed Mediator specifically associating with Pol III in Saccharomyces cerevisiae. Those authors also reported specific associations with RNA polymerase I and proteins involved in transcription elongation and RNA processing, supporting other evidence of Mediator's involvement in elongation and processing.
Chromatin organization
Mediator is involved in chromatin looping, which brings distant regions of a chromosome into closer physical proximity. The ncRNA-a mentioned above is involved in such looping. Enhancer RNAs (eRNAs) can function similarly.
In addition to euchromatin looping, Mediator helps form or maintain heterochromatin at centromeres and telomeres.
Signal transduction
TGFβ signaling at the cell membrane involves two different intracellular pathways. Only one depends on MED15. In both human cells and Caenorhabditis elegans, MED15 helps lipid homeostasis through the SREBP-containing pathway. In the model plant Arabidopsis thaliana, the ortholog of MED15 is required for signaling by the plant hormone salicylic acid, while MED25 is required for the transcriptional activation of responses to hypoxia, jasmonate and shade signalling. Two components of the CDK module (MED12 and MED13) are involved in the Wnt signaling pathway. MED23 is involved in the RAS/MAPK/ERK pathway. This abbreviated review shows the versatility of individual Mediator subunits, and leads to the idea that Mediator is an end-point of signaling pathways.
Human disease
Involvement of Mediator in various human diseases has been reviewed. Since inhibiting one interaction of a disease-causing signaling pathway with a subunit of Mediator may not inhibit general transcription needed for normal function, Mediator subunits are attractive candidates for therapeutic drugs.
Interactions

Very gentle cell lysis in yeast followed by co-immunoprecipitation with an antibody to a MED17 has confirmed almost all previously reported or predicted interactions and revealed many previously unsuspected specific interactions of various proteins with Mediator.
MED1
Main article: MED1

Details of the first subunit are illustrative of the types of information that may be gathered for other subunits. See for them.
Regulation by MicroRNAs
MicroRNAs help regulate the expression of many proteins. MED1 is targeted by miR-1, which is important in gene regulation in cancers. The tumor suppressor miR-137 also regulates MED1.
Mouse embryonic development
Null mutants die early (embryonic day 11.5). Investigating hypomorphic mutants (which survive 2 days longer) found that placental defects were primarily lethal and that there were also defects in cardiac and hepatic development, but many other organs were normal.
Mouse cells and tissues

In mice, conditional mutations can be produced to affect only specific cells or tissues at specific times, so that the mouse can develop to adulthood to have its adult phenotype studied. In one case, MED1 was found to participate in controlling the timing of events of meiosis in male mice. Conditional mutants in keratinocytes differ in skin wound healing. A conditional mutation in mice changed dental epithelium into epidermal epithelium, which caused hair to grow beside the incisors.
Subunit composition
The Mediator complex is composed of at least 31 subunits in all eukaryotes studied: MED1, MED4, MED6, MED7, MED8, MED9, MED10, MED11, MED12, MED13, MED13L, MED14, MED15, MED16, MED17, MED18, MED19, MED20, MED21, MED22, MED23, MED24, MED25, MED26, MED27, MED28, MED29, MED30, MED31, CCNC, and CDK8. There are three fungal-specific components, referred to as MED2, MED3 and MED5.
The subunits form at least three structurally distinct submodules. The head and the middle modules interact directly with RNA polymerase II, whereas the elongated tail module interacts with gene-specific regulatory proteins. Mediator containing the CDK8 module is less active than Mediator lacking this module in supporting transcriptional activation.
- The head module contains: MED6, MED8, MED11, SRB4/MED17, SRB5/MED18, ROX3/MED19, SRB2/MED20 and SRB6/MED22.
- The middle module contains: MED1, MED4, NUT1/MED5, MED7, CSE2/MED9, NUT2/MED10, SRB7/MED21 and SOH1/MED31. CSE2/MED9 interacts directly with MED4.
- The tail module contains: MED2, PGD1/MED3, RGR1/MED14, GAL11/MED15 and SIN4/MED16.
- The CDK8 module contains: MED12, MED13, CCNC and CDK8. Individual preparations of the Mediator complex lacking one or more distinct subunits have been variously termed ARC, CRSP, DRIP, PC2, SMCC and TRAP.
In other species
Below is a cross-species comparison of Mediator complex subunits.
| Subunit No. | Human gene | C. elegans gene | D. melanogaster gene | S. cerevisiae gene | Sch. pombe gene |
|---|---|---|---|---|---|
| MED1 | MED1 | Sop3/mdt-1.1, 1.2 | MED1 | MED1 | med1 |
| MED2 | MED2 | ||||
| MED3 | PGD1 | ||||
| MED4 | MED4 | MED4 | MED4 | med4 | |
| MED5 | NUT1 | ||||
| MED6 | MED6 | MDT-6 | MED6 | MED6 | med6 |
| MED7 | MED7 | MDT-7/let-49 | MED7 | MED7 | med7 |
| MED8 | MED8 | MDT-8 | MED8 | MED8 | med8 |
| MED9 | MED9 | MED9 | CSE2 | ||
| MED10 | MED10 | MDT-10 | NUT2 | med10 | |
| MED11 | MED11 | MDT-11 | MED11 | MED11 | med11 |
| MED12 | MED12 | MDT-12/dpy-22 | MED12 | SRB8 | srb8 |
| MED12L | MED12L | ||||
| MED13 | MED13 | MDT-13/let-19 | MED13 | SSN2 | srb9 |
| MED14 | MED14 | MDT-14/rgr-1 | MED14 | RGR1 | med14 |
| MED15 | MED15 | mdt-15 | MED15 | GAL11 | YN91_SCHPO |
| MED16 | MED16 | MED16 | SIN4 | ||
| MED17 | MED17 | MDT-17 | MED17 | SRB4 | med17 |
| MED18 | MED18 | MDT-18 | MED18 | SRB5 | med18 |
| MED19 | MED19 | MDT-19 | MED19 | ROX3 | med19 |
| MED20 | MED20 | MDT-20 | MED20 | SRB2 | med20 |
| MED21 | MED21 | MDT-21 | MED21 | SRB7 | med21 |
| MED22 | MED22 | MDT-22 | MED22 | SRB6 | med22 |
| MED23 | MED23 | MDT-23/sur-2 | MED23 | ||
| MED24 | MED24 | MED24 | |||
| MED25 | MED25 | MED25 | |||
| MED26 | MED26 | MED26 | |||
| MED27 | MED27 | MED27 | med27 | ||
| MED28 | MED28 | MED28 | |||
| MED29 | MED29 | MDT-19 | MED29 | ||
| MED30 | MED30 | MED30 | |||
| MED31 | MED31 | MDT-31 | MED31 | SOH1 | med31 |
| CCNC | CCNC | cic-1 | CycC | SSN8 | pch1 |
| CDK8 | CDK8 | cdk-8 | Cdk8 | SSN3 | srb10 |
Notes
References
References
- (June 1990). "A novel mediator between activator proteins and the RNA polymerase II transcription apparatus". Cell.
- (April 1991). "A mediator required for activation of RNA polymerase II transcription in vitro". Nature.
- (March 2015). "The Mediator complex: a central integrator of transcription". Nature Reviews Molecular Cell Biology.
- (September 2015). "Molecular architecture of the yeast Mediator complex". eLife.
- (March 2011). "Molecular architecture of the human Mediator-RNA polymerase II-TFIIF assembly". PLOS Biology.
- (3 November 2016). "Mediator Undergoes a Compositional Change during Transcriptional Activation.". Molecular Cell.
- (September 2005). "Yeast mediator and its role in transcriptional regulation". Comptes Rendus Biologies.
- (May 2005). "The yeast Mediator complex and its regulation". Trends in Biochemical Sciences.
- (2015-09-24). "Molecular architecture of the yeast Mediator complex". eLife.
- Soutourina, Julie. (2017-12-06). "Transcription regulation by the Mediator complex". Nature Reviews Molecular Cell Biology.
- (February 2015). "Architecture of the RNA polymerase II–Mediator core initiation complex". Nature.
- (2017). "Chromatin potentiates transcription". Proc Natl Acad Sci U S A.
- (14 January 2016). "The Molecular Basis of Eukaryotic Transcription". Israel Institute for Advanced Studies.
- (February 2016). "Loss of the Mediator subunit Med20 affects transcription of tRNA and other non-coding RNA genes in fission yeast". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms.
- (2017). "Proteomic Analysis of the Mediator Complex Interactome in Saccharomyces cerevisiae". Sci Rep.
- (2002). "A component of the ARC/Mediator complex required for TGF beta/Nodal signalling". Nature.
- (2006). "An ARC/Mediator subunit required for SREBP control of cholesterol and lipid homeostasis". Nature.
- (2012). "Non-recognition-of-BTH4, an Arabidopsis mediator subunit homolog, is necessary for development and response to salicylic acid". Plant Cell.
- (July 2012). "The Arabidopsis Mediator Subunit MED25 Differentially Regulates Jasmonate and Abscisic Acid Signaling through Interacting with the MYC2 and ABI5 Transcription Factors". The Plant Cell.
- (November 2020). "Mediator Subunit MED25 Physically Interacts with PHYTOCHROME INTERACTING FACTOR4 to Regulate Shade-Induced Hypocotyl Elongation in Tomato". Plant Physiology.
- (November 2020). "MED25 Mediates Shade-Induced Hypocotyl Elongation in Tomato". Plant Physiology.
- (11 September 2024). "ERFVII -controlled hypoxia responses are in part facilitated by MEDIATOR SUBUNIT 25 in Arabidopsis thaliana". The Plant Journal.
- (2015). "Mediator kinase module and human tumorigenesis". Crit Rev Biochem Mol Biol.
- (2015). "MED12 and uterine smooth muscle oncogenesis: State of the art and perspectives". Eur J Cancer.
- (2014). "Involvement of Mediator complex in malignancy". Biochim Biophys Acta.
- (2014). "The roles of mediator complex in cardiovascular diseases". Biochim Biophys Acta.
- (2014). "Impaired development of neural-crest cell-derived organs and intellectual disability caused by MED13L haploinsufficiency". Hum Mutat.
- (2013). "Mediator complex dependent regulation of cardiac development and disease". Genomics Proteomics Bioinformatics.
- (2013). "Mediating Lipid Biosynthesis: Implications for Cardiovascular Disease". Trends Cardiovasc Med.
- (2012). "Unraveling framework of the ancestral Mediator complex in human diseases". Biochimie.
- (2011). "Dysregulation of CDK8 and Cyclin C in tumorigenesis". J Genet Genomics.
- (2011). "Mediator and human disease". Semin Cell Dev Biol.
- (2008). "MED12-Related Disorders". University of Washington, Seattle.
- (2014). "MicroRNA-1 functions as a potential tumor suppressor in osteosarcoma by targeting Med1 and Med31". Oncol Rep.
- (2015). "MiR137 is an androgen regulated repressor of an extended network of transcriptional coregulators". Oncotarget.
- (2000). "Involvement of the TRAP220 component of the TRAP/SMCC coactivator complex in embryonic development and thyroid hormone action". Mol Cell.
- (2003). "The thyroid hormone receptor-associated protein TRAP220 is required at distinct embryonic stages in placental, cardiac, and hepatic development". Mol Endocrinol.
- (2015). "Med1 regulates meiotic progression during spermatogenesis in mice". Reproduction.
- (2014). "Alteration of skin wound healing in keratinocyte-specific mediator complex subunit 1 null mice". PLOS ONE.
- (2014). "Ablation of coactivator Med1 switches the cell fate of dental epithelia to that generating hair". PLOS ONE.
- (2004). "A unified nomenclature for protein subunits of mediator complexes linking transcriptional regulators to RNA polymerase II". Molecular Cell.
- "UniProtKB".
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Mediator (coactivator) — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report