From Surf Wiki (app.surf) — the open knowledge base
Haplogroup T-M184
Human Y-chromosome DNA haplogroup
Human Y-chromosome DNA haplogroup
| Field | Value | |
|---|---|---|
| name | T-M184 | |
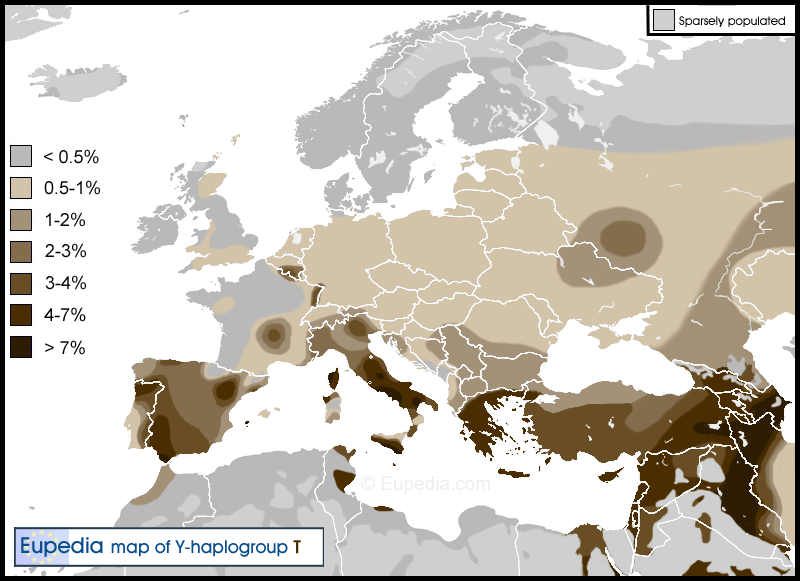
| map | 2026 New Updated Map for Distribution of T-M70.png | |
| origin-date | 26,000 BC BP | |
| origin-place | Between Fertile Crescent & Levant (hypothesized from earliest known sample i1707) | |
| ancestor | LT | |
| descendants | T1 (T-L206); T2 (T-PH110) | |
| mutations | M184/PAGES34/USP9Y+3178, M272, PAGES129, L810, L455, L452, L445 | members = Somali Dir & Isaaq, Fulani (Adamawa & Wodabe), Egyptians, Antemoro (Madagascar); Afar (Ethiopia & Djibouti), Toubou of Chad, Tuareg, Karomojong Nilotes |
| caption | Haplogroup T expansion across the world |
|origin-date = 26,000 BC BP |origin-place= Between Fertile Crescent & Levant (hypothesized from earliest known sample i1707)
Haplogroup T-M184, also known as Haplogroup T, is a human Y-chromosome DNA haplogroup. The unique-event polymorphism that defines this clade is the single-nucleotide polymorphism known as M184.

T-M184 is unusual in that it is both geographically widespread and relatively rare. T1 (T-L206) – the numerically dominant primary branch of T-M184 – appears to have originated in Western Asia, and spread from there into East Africa, South Asia, Europe, Egypt and adjoining regions. T1* may have expanded with the Pre-Pottery Neolithic B culture (PPNB) which originated in West Asia.
The earliest presence of T-M184 appears in Ain Ghazal, Jordan (sample i1707), bordering Asia and Africa. The individual predated the arrival of Caucaso-Iranian ancestry to the Levant. His DNA consisted of Natufian Hunter Gatherer and Anatolian Neolithic ancestry, together known as PPNB, which was the indigenous ancestry of the Levant at the time.
Subclades of T-M70 appear to have been present in Europe since the Neolithic with Neolithic Farmers from Western Asia. The moderately high frequency (~18%) of T1b* chromosomes in the Lemba of southern Africa supports the hypothesis of a West Asian origin for their paternal line.
Structure
Haplogroup_K = ~45,000–50,000 years ago",
"Region": "South or Southeast Asia"
"Descendants" "LT (K1)"~40,000–45,000 years ago",
"Region": "South Asia / Iranian Plateau", "
Descendants": L~25,000–30,000 years ago",
"Region": "South Asia, Dravidians, Indus Valley",
"Notes": "Possibly linked to pre-Neolithic South Asian populations
| "T": ~25,000–30,000 years ago", |
"Region": "South Asia,",
"Notes": "Later spread into the Levant, Mediterranean, and Africa" ;Subclade structure of Haplogroup T (M184).
- T1 (L206)
- T1a (M70/Page46/PF5662)
- T1a1 (L162/Page21, L454)
- T1a1a (L208/Page2)
- T1a1a1 (CTS11451)
- T1a1a2 (Y16897)
- T1a1a2a (Z19963)
- T1a1a (L208/Page2)
- T1a2 (L131)
- T1a2a (PH141/Y13244)
- T1a2b (L446)
- T1a3 (FGC1350/Y11151 )
- T1a3a (Y11675/Z9798)
- T1a3b (FGC1340/Y8614)
- T1a1 (L162/Page21, L454)
- T1a (M70/Page46/PF5662)
- T2 (PH110)
Distribution
Overview
As a primary branch of haplogroup LT (a.k.a. K1), the basal, undivergent haplogroup T* currently has the alternate phylogenetic name of K1b and is a sibling of haplogroup L* (a.k.a. K1a). (Before 2008, haplogroup T and its subclades were known as haplogroup K2. The name K2 has since been reassigned to a primary subclade of haplogroup K.) It has two primary branches: T1 (T-L206) and T2 (T-PH110). Most males who now belong to haplogroup T1* carry the subclade T-M70 (T1a), a primary branch of T-M206.
Haplogroup T is found at exceptionally high levels amongst the Dir and Isaaq in Somaliland, Djibouti, and Ethiopia. it is also found at relatively high levels in specific populations in other parts of the world especially amongst Arabs from UAE in South Eastern Arabia T-M184 spikes at 19% on FTDNA. These include Kurru, Bauris and Lodha in South Asia; among Toubou in Chad; Somalilander clans: Isaaq and Dir, southern Egyptians and Fula (Fulbe) in north Cameroon; ; Zoroastrians, Bakhtiaris, Assyrians and Iraqi Jews in the Middle East. T is a rather rare haplogroup, displaying a global frequency of around 1% (King et al., 2007), but nonetheless it is found at quite high frequencies in Sephardic Levites (23%) and Sephardic Israelis (13%; Behar et al., 2004).
The maximal worldwide frequency for haplogroup T-M184 is 100%, amongst Dir clan males (Iacovacci et al. 2016). [6] It accounts for approximately 82.4% of ethnic Somali male lineages overall in Dire Dawa, Ethiopia (Plaster et al. 2011). T is only 9% in Somalia (Iacovacci et al. 2016). Geographically, it is found at the highest levels in the Dire Dawa area of Ethiopia, and Djibouti.
Luis et al. (2004) suggest that the presence of T on the African continent may, like R1* representatives, point to an older introduction from West Asia. The Levant rather than the Arabian Peninsula appears to have been the main route of entry, as the Egyptian and Anatolian haplotypes are considerably older in age (13,700 BP and 9,000 BP, respectively) than those found in Oman (only 1,600 BP). According to the authors, haplogroup T-M184 within Africa represents the traces of a more widespread early local presence of the West Asian clade. Later expansions of populations from West Asia carrying the E-M215, E-V38, G and J NRY lineages may have overwhelmed the T-M184 clade-bearers in certain localities.
In the Caucasus and Anatolia it makes up to 4% of the population in southeast and northwest Caucasus as well as in southeast and western Anatolia, peaking up to 20% in Armenians from Sasun. In Middle East it makes up to 4% of the population around the Zagros Mountains and the Persian Gulf as well as around the Taurus Mountains and the Levant basin, peaking up to 10% in Zoroastrians from Kerman, Bakhtiaris, Assyrians (up to 40%), Abudhabians, Armenians from Historical Southwestern Armenia and Druzes from Galilee. In Eastern Africa, it makes up to 4% of the population on Upper Egypt peaking up to 10% in Luxor.
Haplogroup T is uncommon in Europe, except in Southern Europe and adjoining areas. According to Mendez et al. (2011), "the occurrence in Europe of lineages belonging to both T1a1 (old T1a) and T1a2 (old T1b) subclades probably reflects multiple episodes of gene flow. T1a1* haplogroups in Europe likely reflect older gene flow". It makes up to 4% of the population in Central Italy, Western Sicily, Northwest Corsica, Northwestern Iberian Peninsula, Western Andalucia, Western Alps, Eastern Crete, and Macedonia, frequencies up to 10% in Ibiza, Miranda de I Douro, Eastern Oviedo, Cádiz, Badajoz, Balagna, Norma and Ragusa, and peaking at 20% in Sciacca, L'Aquila and some German southern regions. T-M184 was found in 1.7% (10/591) of a pool of six samples of males from southwestern Russia, but it was completely absent from a pool of eight samples totalling 637 individuals from the northern half of European Russia. The Russians from the southwest were from the following cities: Roslavl, Livny, Pristen, Repyevka, and Belgorod; and Kuban Cossacks from the Republic of Adygea.
T1 (T*)
| Macedonians | Macedonian | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| (Balto-Slavic) | Macedonia | 1/201 | 0.5% | vauthors=Jakovski Z, Nikolova K, Jankova-Ajanovska R, Marjanovic D, Pojskic N, Janeska B | title=Genetic data for 17 Y-chromosomal STR loci in Macedonians in the Republic of Macedonia | journal=Forensic Science International. Genetics | volume=5 | issue=4 | pages=e108–11 | year=2011 | pmid=21549657 | doi=10.1016/j.fsigen.2011.04.005}} | Macedonians Orthodox Christians |
Main article: Haplogroup T-L206
T1 is the most common descent of T-M184 haplogroup, being the lineage of more than 95% of all Eurasian T-M184 members. One of their descent lineages is found in high frequencies among northern Somali clans. However, it appears to have originated somewhere around the Eastern Mediterranean Basin, perhaps somewhere between Palestine to the Jordan Valley.
The basal T1* subclade appears to have spread to northeastern Anatolia, from the Levant and Mesopotamia at least, with the Pre-Pottery Neolithic B culture (PPNB). Although it is rare in modern populations, T1* has been found in a Berber individual from Tunisia, a male in Syria, and one sequence among ethnic Macedonians in Macedonia.
T1a (M70)
Mendez et al. (2011) points to an ancient presence for T1a-M70 in Europe may reflect early exiles between the ancient lands of Israel and Babylonia and Assyria. The subclade probably arrived with the very first farmers.
T1a1*
With the comparison of the data provided by the Pityusic population with other circumediterranean populations surprises that practically there is no convergence with any of these populations, not even with the North African populations. The Pityusic case is paradigmatic: for some markers shows affinities with Oriental populations (some mtDNA variables), but diverges from these populations when considering other markers. It is a separate case, an island, not in the geographical sense but genetical.
The Pityusans of the Pityusic Islands (Ibiza and Formentera) – have been found by three different studies to possess T1a1 at relatively high levels of 6.7–16.7%. Tomàs et al. (2006) found three cases amongst a sample of 45 (6.7%). Zalloua et al. (2008) found nine examples that were L454+ (an SNP equivalent to L162/Page21) from a sample of 54 (i.e. a rate of 16.7%). Rodriguez et al. (2009) found seven cases of L454+ in a sample of 96 (7.3%).
The Pontic Greeks of Anatolia are also reported to possess T1a1. In 2009, a male with the surname Metaxopoulos and a Pontic Greek background was reported to be T-L162(xL208) – according to the Y-Chromosome Genome Comparison Project administered by Adriano Squecco. Greeks from the Fatsa (originally "Φάτσα") reportedly migrated in antiquity from Sinope, which was itself colonised by Ionians (from Miletus). Another ancient Ionian colony in north-west Anatolia, Lámpsakos (Lampsacus), had onomastic links to the Pityusic Islands (see above) – Lámpsakos was originally an Ionian colony known as Pityussa.
T1a1a (L208)
This lineage, formed 14,200-11,000 BP, is the largest branch downstream T1a1-L162. The Isaaq clans and dir is T-L208, they found in Somaliland, Djibouti, Somalia and Somali Ethiopia.
T1a1a1a1b1a1* (T-Y3782*)
One Sardinian male from a sample of 187 (a nominal rate of 0.53%) – a resident of the Province of Cagliari (Sardinian: Casteddu) – has been found to have T-Y3782(xY3836), also known T1a1a1a1b1a1(xT1a1a1a1b1a1a).
T1a1a1a1b1a1a (T-Y3836)

This lineage is mostly found among individuals from the Iberian Peninsula, where the subclade also has its highest diversity. Two subclades can be clearly discriminated. The first, found mainly in post-colonial Puerto Rico, with DYS391=10 and the second, found mainly in Panamá where their Iberian descendants could have the entrance point to America, with DYS439=12.
Some members of Y3836 are found among different communities of the Sephardic diaspora but they are found to be extremely rare in the total percentage of some of these communities as seen in Nogueiro et al. This probably could mean that these members could be integrated by these communities through the contact with other native Iberian populations as seen in Monteiro et al. where this lineage was found among native Astur-Leonese speakers.
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Panamanians | Panamian Castilian (Romance languages) | Los Santos Province | 1/30 | 3.3% | ||||||||||||
| Colombians | Colombian Castilian (Romance languages) | Caldas | 2/75 | 2.7% | YHRD | Mestizo individuals | ||||||||||
| Panamanians | Panamian Castilian (Romance languages) | Panama Province | 1/43 | 2.3% | ||||||||||||
| Northwest Argentinians | Argentinian Castilian (Romance languages) | Mountainous region of Jujuy | 1/50 | 2% | YHRD | Admixed population | ||||||||||
| Puerto Ricans | Puerto Rican Castilian (Romance languages) | Southeast Puerto Rico | 2/110 | 1.8% | vauthors=Vilar MG, Melendez C, Sanders AB, Walia A, Gaieski JB, Owings AC, Schurr TG | title=Genetic diversity in Puerto Rico and its implications for the peopling of the Island and the West Indies | journal=American Journal of Physical Anthropology | volume=155 | issue=3 | pages=352–68 | year=2014 | pmid=25043798 | doi=10.1002/ajpa.22569 | bibcode=2014AJPA..155..352V | s2cid=205334949 }} | |
| Northeastern Portuguese Jews | Judaeo-Portuguese (Romance) | Bragança, Argozelo, Carção, Mogadouro, and Vilarinho dos Galegos | 1/57 | 1.8% | ||||||||||||
| Native Mirandese speakers | Mirandese Astur-Leonese (Romance) | Miranda de l Douro | 1/58 | 1.7% | ||||||||||||
| Dominicans | Dominican Castilian (Romance languages) | Dominican Republic | 4/261 | 1.5% | vauthors=Díaz V, Carracedo A | title=The distribution of Y-chromosome STRs in Dominican population | journal=Forensic Science International: Genetics Supplement Series | volume=1 | issue=1 | year=2008 | pages=195–7 | doi=10.1016/j.fsigss.2007.10.163 | doi-access=free }} | |||
| Panamanians | Panamian Castilian (Romance languages) | Chiriquí Province | 1/92 | 1.1% | ||||||||||||
| Mecklenburgers | East Low Saxon (West Germanic) | Rostock | 2/200 | 1% | ||||||||||||
| Mestizos | Colombian Castilian (Romance languages) | Bogotá | 2/195 | 1% | YHRD | |||||||||||
| Mestizos | Colombian Castilian (Romance languages) | Valle del Cauca | 1/103 | 1% | YHRD | |||||||||||
| Mestizos | Ecuadorian Castilian (Romance languages) | Quito | 1/102 | 1% | vauthors=González-Andrade F, Roewer L, Willuweit S, Sánchez D, Martínez-Jarreta B | title=Y-STR variation among ethnic groups from Ecuador: Mestizos, Kichwas, Afro-Ecuadorians and Waoranis | journal=Forensic Science International. Genetics | volume=3 | issue=3 | pages=e83–91 | year=2009 | pmid=19414158 | doi=10.1016/j.fsigen.2008.08.003}} | |||
| Venezuelans | Venezuelan Castilian (Romance languages) | Maracaibo | 1/111 | 0.9% | vauthors=Borjas L, Bernal LP, Chiurillo MA, Tovar F, Zabala W, Lander N, Ramírez JL | title=Usefulness of 12 Y-STRs for forensic genetics evaluation in two populations from Venezuela | journal=Legal Medicine | volume=10 | issue=2 | pages=107–12 | year=2008 | pmid=17981491 | doi=10.1016/j.legalmed.2007.08.005}} | |||
| Venezuelans | Venezuelan Castilian (Romance languages) | Central Region | 1/115 | 0.9% | vauthors=Alvarez M, Marrero C, Dictamen A, Figuera M, Marrero M, Borjas L, Ferreira R | title=Y-chromosome haplotype database in Venezuelan central region and its comparison with other Venezuelan populations | journal=Forensic Science International: Genetics Supplement Series | volume=2 | issue=1 | year=2009 | pages=407–8 | doi=10.1016/j.fsigss.2009.08.100}} | ||||
| Europeans | Brazilian Portuguese (Romance languages) | São Paulo | 1/120 | 0.8 | YHRD | European descents | ||||||||||
| Ecuadorians | Ecuadorian Castilian (Romance languages) | Quito | 1/120 | 0.8% | ||||||||||||
| Colombians | Colombian Castilian (Romance languages) | Antioquia | 6/777 | 0.7% | vauthors=Builes JJ, Bravo ML, Gómez C, Espinal C, Aguirre D, Gómez A, Rodríguez J, Castañeda P, Montoya A, Moreno M, Amorim A, Gusmão L | title=Y-chromosome STRs in an Antioquian (Colombia) population sample | journal=Forensic Science International | volume=164 | issue=1 | pages=79–86 | year=2006 | pmid=16289613 | doi=10.1016/j.forsciint.2005.10.005}} | |||
| Mexicans | Mexican Castilian (Romance languages) | Mérida | 1/159 | 0.6% | YHRD | Mestizo individuals | ||||||||||
| Eastern Andalusians | Andalusian (Romance) | Alhama de Granada, Baza, Huéscar, Loja, Montefrío and Órgiva | 1/180 | 0.6% | ||||||||||||
| Colombians | Colombian Castilian (Romance languages) | Santander | 1/193 | 0.5% | YHRD | Mestizo individuals | ||||||||||
| Chileans | Chilean Castilian (Romance languages) | Concepción | 1/198 | 0.5% | YHRD | |||||||||||
| Catalans | Not reported | Metropolitan area of Barcelona | 1/224 | 0.5% | vauthors=Gené M, Borrego N, Xifró A, Piqué E, Moreno P, Huguet E | title=Haplotype frequencies of eight Y-chromosome STR loci in Barcelona (North-East Spain) | journal=International Journal of Legal Medicine | volume=112 | issue=6 | pages=403–5 | year=1999 | pmid=10550606 | doi=10.1007/s004140050025 | s2cid=29850287 }} | ||
| Mexicans | Mexican Spanish (Romance languages) | Guadalajara | 1/246 | 0.4% | YHRD | Mestizo individuals | ||||||||||
| Europeans | Brazilian Portuguese (Romance languages) | Rio Grande do Sul | 1/255 | 0.4% | vauthors=Schwengber SP, Kommers T, Matte CH, Raimann PE, Carvalho BA, Leite FP, Medeiros MA, Souza LF, Castro CS, Chassot FG, Bonatto SL | title=Population data of 17 Y-STR loci from Rio Grande do Sul state (South Brazil) | journal=Forensic Science International. Genetics | volume=4 | issue=1 | pages=e31–3 | year=2009 | pmid=19948319 | doi=10.1016/j.fsigen.2009.02.001}} |
T2 (PH110)
This lineage could have arrived in the Levant through the PPNB expansion from northeastern Anatolia.
A 2014 study found T-PH110 in one ethnic Bhutanese male, out of a sample of 21, possibly implying a rate of 4.8% in Bhutan. Also have been found in a German individual and another two from Caucasus. The Bhutanese and the German haplotypes seems to cluster together.
Possible cases from older research
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Altaians | Altai (Turkic) | Kurmach-Baygol | 2/11 | 18.2% | vauthors=Khar'kov VN, Stepanov VA, Medvedeva OF, Spiridonova MG, Voevoda MI, Tadinova VN, Puzyrev VP | title=[Gene pool differences between northern and southern Altaians inferred from the data on Y-chromosomal haplogroups] | language=ru | journal=Genetika | volume=43 | issue=5 | pages=675–87 | year=2007 | pmid=17633562 | doi=10.1134/S1022795407050110 | s2cid=566825 }} | K* (xT1a-M70, L-M20, N-DYF155S2, O-M175, P-92R7) | |
| Altaians | Altai (Turkic) | Turochak | 2/19 | 10.5% | K(xT1a-M70, L-M20, N-DYF155S2, O-M175, P-92R7) | ||||||||||||
| Leoneses | Astur-Leonese (Romance) | Leon | 1/13 | 7.7% | K(xT1a-M70, L1-M22, P-92R7) | ||||||||||||
| Ossetian Irons | Iron (Iranian) | South Ossetia | 1/21 | 4.8% | vauthors=Bekada A, Fregel R, Cabrera VM, Larruga JM, Pestano J, Benhamamouch S, González AM | title=Introducing the Algerian mitochondrial DNA and Y-chromosome profiles into the North African landscape | journal=PLOS ONE | volume=8 | issue=2 | article-number=e56775 | year=2013 | pmid=23431392 | pmc=3576335 | doi=10.1371/journal.pone.0056775 | bibcode=2013PLoSO...856775B | doi-access=free }} | No further details available. |
| Cordobeses | Andalusian (Romance) | Córdoba | 1/27 | 3.7% | vauthors=López-Parra AM, Gusmão L, Tavares L, Baeza C, Amorim A, Mesa MS, Prata MJ, Arroyo-Pardo E | title=In search of the pre- and post-neolithic genetic substrates in Iberia: evidence from Y-chromosome in Pyrenean populations | journal=Annals of Human Genetics | volume=73 | issue=1 | pages=42–53 | year=2009 | pmid=18803634 | doi=10.1111/j.1469-1809.2008.00478.x | s2cid=43273988 }} | No further details available. | ||
| Leoneses | Astur-Leonese (Romance) | Leon | 2/60 | 3.3% | No further details available. | ||||||||||||
| Tharus | Tharu (Indo-Aryan) | Morang | 1/37 | 2.7% | vauthors=Fornarino S, Pala M, Battaglia V, Maranta R, Achilli A, Modiano G, Torroni A, Semino O, Santachiara-Benerecetti SA | title=Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation | journal=BMC Evolutionary Biology | volume=9 | article-number=154 | year=2009 | issue=1 | pmid=19573232 | pmc=2720951 | doi=10.1186/1471-2148-9-154 | bibcode=2009BMCEE...9..154F | doi-access=free }} | K(xT1a-M70, L-M20, NO-M214, P-M74) |
| Cherkessians | Besleney (Northwest Caucasian) | Circassia | 2/126 | 1.6% | No further details are available. | ||||||||||||
| Bizkaians | Bizkaiera (Isolate language) | Bizkaia | 1/72 | 1.4% | No further details are available. | ||||||||||||
| Europeans | English (Germanic) | Australia | 1/1078 | 0.09% | vauthors=Taylor DA, Henry JM | title=Haplotype data for 16 Y-chromosome STR loci in Aboriginal and Caucasian populations in South Australia | journal=Forensic Science International. Genetics | volume=6 | issue=6 | pages=e187–8 | year=2012 | pmid=22673611 | doi=10.1016/j.fsigen.2012.05.005}} | No further details are available. |
Modern geographical distribution
Northern Asia
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Kazakhs | Kazakh (Turkic) | Southwestern Altai | 1/30 | 3.3% | T1a-M70 | |||||||||||
| Evens | Even (Tungusic) | eastern Siberia | 1/61 | 1.6% | ||||||||||||
| Barghuts | Barga (Mongolic) | different localities of Hulun Buir Aimak | 1/76 | 1.3% | vauthors=Malyarchuk BA, Derenko M, Denisova G, Woźniak M, Rogalla U, Dambueva I, Grzybowski T | title=Y chromosome haplotype diversity in Mongolic-speaking populations and gene conversion at the duplicated STR DYS385a,b in haplogroup C3-M407 | journal=Journal of Human Genetics | volume=61 | issue=6 | pages=491–6 | year=2016 | pmid=26911356 | doi=10.1038/jhg.2016.14 | s2cid=13217444 | doi-access=free }} | T1a-M70. In the 12–13th centuries, the Barga (Barghuts) Mongols appeared as tribes near Lake Baikal, named Bargujin. |
Europe
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Marchigianos | Marchigiano dialect (Italian) | Arquata del Tronto and Apiro | 2/2 | 100% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cretans and southern Aegeans | Southeastern Greek | Crete and southern Aegean | 2/6 | 33.3% | vauthors=Katsaloulis P, Tsekoura K, Vouropoulou M, Miniati P | title=Genetic population study of 11 Y chromosome STR loci in Greece | journal=Forensic Science International. Genetics | volume=7 | issue=3 | pages=e56–8 | year=2013 | pmid=23582698 | doi=10.1016/j.fsigen.2013.02.001}} | |||||||||||||||||||||||||||||||||||||
| Rural Saccensi | Sicilian (Romance) | Sciacca | 6/20 | 30% | vauthors=Robino C, Ralf A, Pasino S, De Marchi MR, Ballantyne KN, Barbaro A, Bini C, Carnevali E, Casarino L, Di Gaetano C, Fabbri M, Ferri G, Giardina E, Gonzalez A, Matullo G, Nutini AL, Onofri V, Piccinini A, Piglionica M, Ponzano E, Previderè C, Resta N, Scarnicci F, Seidita G, Sorçaburu-Cigliero S, Turrina S, Verzeletti A, Kayser M | title=Development of an Italian RM Y-STR haplotype database: Results of the 2013 GEFI collaborative exercise | journal=Forensic Science International. Genetics | volume=15 | pages=56–63 | year=2015 | pmid=25457630 | doi=10.1016/j.fsigen.2014.10.008 | hdl=2318/154001 | hdl-access=free }} | ||||||||||||||||||||||||||||||||||||
| Chians | Southeastern Greek | Khíos | 4/16 | 25% | vauthors=Robino C, Varacalli S, Gino S, Chatzikyriakidou A, Kouvatsi A, Triantaphyllidis C, Di Gaetano C, Crobu F, Matullo G, Piazza A, Torre C | title=Y-chromosomal STR haplotypes in a population sample from continental Greece, and the islands of Crete and Chios | journal=Forensic Science International | volume=145 | issue=1 | pages=61–4 | year=2004 | pmid=15374596 | doi=10.1016/j.forsciint.2004.02.026}} | |||||||||||||||||||||||||||||||||||||
| Stilfser (Tyrolese) | Southern Austro-Bavarian (German) | Stilfs, South Tyrol, Italy | 4/17 | 23.5% | vauthors=Pichler I, Mueller JC, Stefanov SA, De Grandi A, Volpato CB, Pinggera GK, Mayr A, Ogriseg M, Ploner F, Meitinger T, Pramstaller PP | title=Genetic structure in contemporary south Tyrolean isolated populations revealed by analysis of Y-chromosome, mtDNA, and Alu polymorphisms | journal=Human Biology | volume=78 | issue=4 | pages=441–64 | year=2006 | pmid=17278620 | doi=10.1353/hub.2006.0057 | s2cid=20205296 }} | ||||||||||||||||||||||||||||||||||||
| Sephardic Levites | 7/31 | 22.6% | vauthors=Behar DM, Thomas MG, Skorecki K, Hammer MF, Bulygina E, Rosengarten D, Jones AL, Held K, Moses V, Goldstein D, Bradman N, Weale ME | title=Multiple origins of Ashkenazi Levites: Y chromosome evidence for both Near Eastern and European ancestries | journal=American Journal of Human Genetics | volume=73 | issue=4 | pages=768–79 | year=2003 | pmid=13680527 | pmc=1180600 | doi=10.1086/378506}} | Among Ashkenazi Levites found at 3.3% but different haplotype. | |||||||||||||||||||||||||||||||||||||
| Venetians | Venetian (Romance) | Vigasio and Povegliano Veronese | 2/9 | 22.2% | vauthors=Turrina S, Atzei R, De Leo D | title=Y-chromosomal STR haplotypes in a Northeast Italian population sample using 17plex loci PCR assay | journal=International Journal of Legal Medicine | volume=120 | issue=1 | pages=56–9 | year=2006 | pmid=16328424 | doi=10.1007/s00414-005-0054-x | s2cid=237262 }} | ||||||||||||||||||||||||||||||||||||
| Abruzzesi | Neapolitan language (Romance) | L'Aquila | 6/30 | 20% | vauthors=Boattini A, Martinez-Cruz B, Sarno S, Harmant C, Useli A, Sanz P, Yang-Yao D, Manry J, Ciani G, Luiselli D, Quintana-Murci L, Comas D, Pettener D | title=Uniparental markers in Italy reveal a sex-biased genetic structure and different historical strata | journal=PLOS ONE | volume=8 | issue=5 | article-number=e65441 | year=2013 | pmid=23734255 | pmc=3666984 | doi=10.1371/journal.pone.0065441 | bibcode=2013PLoSO...865441B | doi-access=free }} | macro-haplogroup LT is 30% in L'Aquila population. This was the land of Samnium inhabited by the Caraceni | |||||||||||||||||||||||||||||||||
| Cretans | Cretan Greek | Lasithi | 9/50 | 18% | According to Martinez2007 only can belong to T1a-M70 | |||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Sciacca | 5/28 | 17.9% | vauthors=Di Gaetano C, Cerutti N, Crobu F, Robino C, Inturri S, Gino S, Guarrera S, Underhill PA, King RJ, Romano V, Cali F, Gasparini M, Matullo G, Salerno A, Torre C, Piazza A | title=Differential Greek and northern African migrations to Sicily are supported by genetic evidence from the Y chromosome | journal=European Journal of Human Genetics | volume=17 | issue=1 | pages=91–9 | year=2009 | pmid=18685561 | pmc=2985948 | doi=10.1038/ejhg.2008.120}} | ||||||||||||||||||||||||||||||||||||
| Urban Ragusani | Sicilian (Romance) | Ragusa | 3/19 | 15.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northeastern Portuguese Jews | Judaeo-Portuguese (Romance) | Bragança, Argozelo, Carção, Mogadouro, and Vilarinho dos Galegos | 9/57 | 15.7% | vauthors=Nogueiro I, Manco L, Gomes V, Amorim A, Gusmão L | title=Phylogeographic analysis of paternal lineages in NE Portuguese Jewish communities | journal=Am. J. Phys. Anthropol. | volume=141 | issue=3 | pages=373–81 | date=March 2010 | pmid=19918998 | doi=10.1002/ajpa.21154 | bibcode=2010AJPA..141..373N | doi-access=free }} | T have been found to be the second largest lineage in the Mirandês speaking population of Miranda do Douro too. Haplogroup T was not found in a sample of Belmonte Jews. | ||||||||||||||||||||||||||||||||||
| Albanians | Albanian | Brescia (Lombardia) | 12/83 | 14.5% | vauthors=Cortellini V, Verzeletti A, Cerri N, Marino A, De Ferrari F | title=Y-chromosome polymorphisms and ethnic group - a combined STR and SNP approach in a population sample from northern Italy | journal=Croatian Medical Journal | volume=54 | issue=3 | pages=279–85 | year=2013 | pmid=23771759 | pmc=3692336 | doi=10.3325/cmj.2013.54.279}} | The haplogroup tested is K*(xNOP), is assumed as LT and most probably are members of T | |||||||||||||||||||||||||||||||||||
| Rural Normensi | Italian (Romance) | Norma | 1/7 | 14.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Corsicans | Corsican (Romance) | Balagne (region of Corsica suprana) | 3/24 | 12.5% | vauthors=Scozzari R, Cruciani F, Pangrazio A, Santolamazza P, Vona G, Moral P, Latini V, Varesi L, Memmi MM, Romano V, De Leo G, Gennarelli M, Jaruzelska J, Villems R, Parik J, Macaulay V, Torroni A | title=Human Y-chromosome variation in the western Mediterranean area: implications for the peopling of the region | journal=Human Immunology | volume=62 | issue=9 | pages=871–84 | year=2001 | pmid=11543889 | doi=10.1016/S0198-8859(01)00286-5}} | |||||||||||||||||||||||||||||||||||||
| Rural Piazzesi | Sicilian (Romance) | Piazza Armerina | 3/24 | 12.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Frosinonensis | Central Italian language (Romance) | Filettino | 2/17 | 11.8% | vauthors=Messina F, Finocchio A, Rolfo MF, De Angelis F, Rapone C, Coletta M, Martínez-Labarga C, Biondi G, Berti A, Rickards O | title=Traces of forgotten historical events in mountain communities in Central Italy: A genetic insight | journal=American Journal of Human Biology | volume=27 | issue=4 | pages=508–19 | year=2015 | pmid=25728801 | doi=10.1002/ajhb.22677 | s2cid=30111156 }} | Isolated mountain community | |||||||||||||||||||||||||||||||||||
| Vellepetrianis | Central Italian language (Romance) | Vallepietra | 2/18 | 11.1% | Isolated mountain community | |||||||||||||||||||||||||||||||||||||||||||||
| Cantabrians | Astur-Leonese (Romance) | Cantabria | 2/18 | 11.1% | vauthors=Martínez-Cruz B, Harmant C, Platt DE, Haak W, Manry J, Ramos-Luis E, Soria-Hernanz DF, Bauduer F, Salaberria J, Oyharçabal B, Quintana-Murci L, Comas D | title=Evidence of pre-Roman tribal genetic structure in Basques from uniparentally inherited markers | journal=Molecular Biology and Evolution | volume=29 | issue=9 | pages=2211–22 | year=2012 | pmid=22411853 | doi=10.1093/molbev/mss091 | doi-access=free | hdl=10261/112478 | hdl-access=free }} | All individuals were interviewed in order to assess the geographical origin of their grandparents and their speaking dialect. | |||||||||||||||||||||||||||||||||
| Marchigianos | Marchigiano (Romance) | Matelica | 1/9 | 11.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gaditanos | Andalusian (Romance) | Cádiz | 3/28 | 10.7% | vauthors=Flores C, Maca-Meyer N, González AM, Oefner PJ, Shen P, Pérez JA, Rojas A, Larruga JM, Underhill PA | title=Reduced genetic structure of the Iberian peninsula revealed by Y-chromosome analysis: implications for population demography | journal=European Journal of Human Genetics | volume=12 | issue=10 | pages=855–63 | year=2004 | pmid=15280900 | doi=10.1038/sj.ejhg.5201225 | doi-access=free }} | ||||||||||||||||||||||||||||||||||||
| Native Mirandese speakers | Astur-Leonese (Romance) | Miranda de l Douro | 6/58 | 10.4% | hdl=10216/65272 | last1=Monteiro | first1=Sofia Lucília Monteiro Marques | year=2012 | title=Leonese dialects in Portugal: linguistic-genetic relationships through Y chromosome analysis | type=PhD Thesis | publisher=Universidade do Porto | doi=10.34626/5rpg-3p72 }} | ||||||||||||||||||||||||||||||||||||||
| Pacenses | Astur-Leonese (Romance) | Badajoz | 3/29 | 10.3% | vauthors=Martinez-Cadenas C, Blanco-Verea A, Hernando B, Busby GB, Brion M, Carracedo A, Salas A, Capelli C | title=The relationship between surname frequency and Y chromosome variation in Spain | journal=European Journal of Human Genetics | volume=24 | issue=1 | pages=120–8 | year=2016 | pmid=25898922 | pmc=4795233 | doi=10.1038/ejhg.2015.75}} | ||||||||||||||||||||||||||||||||||||
| Asturianos | Astur-Leonese (Romance) | Eastern Uviéu | 1/10 | 10% | vauthors=Pardiñas AF, Roca A, García-Vazquez E, López B | title=Assessing the genetic influence of ancient sociopolitical structure: micro-differentiation patterns in the population of Asturias (Northern Spain) | journal=PLOS ONE | volume=7 | issue=11 | article-number=e50206 | year=2012 | pmid=23209673 | pmc=3507697 | doi=10.1371/journal.pone.0050206 | bibcode=2012PLoSO...750206P | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Murcianos | Murcian (Romance) | Murcia | 1/10 | 10% | vauthors=Santos C, Fregel R, Cabrera VM, Alvarez L, Larruga JM, Ramos A, López MA, Pilar Aluja M, González AM | title=Mitochondrial DNA and Y-chromosome structure at the Mediterranean and Atlantic façades of the Iberian Peninsula | journal=American Journal of Human Biology | volume=26 | issue=2 | pages=130–41 | year=2014 | pmid=24375863 | doi=10.1002/ajhb.22497 | s2cid=205303141 }} | ||||||||||||||||||||||||||||||||||||
| Aquilanis | Neapolitan language (Romance) | Cappadocia | 5/54 | 9.3% | Isolated mountain community | |||||||||||||||||||||||||||||||||||||||||||||
| Rural Alcamesi | Sicilian (Romance) | Alcamo | 2/22 | 9.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cretans | Cretan Greek | Lasithi | 2/23 | 8.7% | vauthors=Martinez L, Underhill PA, Zhivotovsky LA, Gayden T, Moschonas NK, Chow CE, Conti S, Mamolini E, Cavalli-Sforza LL, Herrera RJ | title=Paleolithic Y-haplogroup heritage predominates in a Cretan highland plateau | journal=European Journal of Human Genetics | volume=15 | issue=4 | pages=485–93 | year=2007 | pmid=17264870 | doi=10.1038/sj.ejhg.5201769 | doi-access=free }} | ||||||||||||||||||||||||||||||||||||
| Ligurians and Tuscans | Ligurian (Romance) | La Spezia / Massa | 2/24 | 8.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lugueses | Galician language (Romance) | Lugo | 1/12 | 8.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Campanians | Neapolitan language (Romance) | West Campania | 7/84 | 8.3% | vauthors=Capelli C, Brisighelli F, Scarnicci F, Arredi B, Caglia' A, Vetrugno G, Tofanelli S, Onofri V, Tagliabracci A, Paoli G, Pascali VL | title=Y chromosome genetic variation in the Italian peninsula is clinal and supports an admixture model for the Mesolithic-Neolithic encounter | journal=Molecular Phylogenetics and Evolution | volume=44 | issue=1 | pages=228–39 | year=2007 | pmid=17275346 | doi=10.1016/j.ympev.2006.11.030 | bibcode=2007MolPE..44..228C }} | ||||||||||||||||||||||||||||||||||||
| Campanians | Neapolitan language (Romance) | Cilento | 4/48 | 8.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Alcamo | 2/24 | 8.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lebaniegos | Astur-Leonese (Romance) | Liébana | 3/37 | 8.1% | last1=Maca-Meyer | first1=N. | last2=Sánchez-Velasco | first2=P. | last3=Flores | first3=C. | last4=Larruga | first4=J.-M. | last5=Gonzalez | first5=A.-M. | last6=Oterino | first6=A. | last7=Leyva-Cobian | first7=F. | year=2003 | title=Y Chromosome and Mitochondrial DNA Characterization of Pasiegos, a Human Isolate from Cantabria (Spain) | journal=Annals of Human Genetics | volume=67 | issue=4 | pages=329–339 | doi=10.1046/j.1469-1809.2003.00045.x | pmid=12914567 | s2cid=40355653 }} | |||||||||||||||||||||||
| Corsicans | Corsican (Romance) | Corte (region of Corsica suprana) | 5/62 | 8.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Segovianos | Castilian language (Romance) | Segovia | 2/25 | 8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Marchigianos | Marchigiano (Romance) | Offida | 3/38 | 7.9% | vauthors=Brisighelli F, Blanco-Verea A, Boschi I, Garagnani P, Pascali VL, Carracedo A, Capelli C, Salas A | title=Patterns of Y-STR variation in Italy | journal=Forensic Science International. Genetics | volume=6 | issue=6 | pages=834–9 | year=2012 | pmid=22487686 | doi=10.1016/j.fsigen.2012.03.003 | hdl=20.500.11940/670 | hdl-access=free }} | |||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | East Sicily | 9/114 | 7.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Saracinescanis | Central Italian language (Romance) | Saracinesco | 2/18 | 7.7% | Isolated mountain community | |||||||||||||||||||||||||||||||||||||||||||||
| Croats | Croatian (West Slavic) | Mljet Island | 3/39 | 7.7% | vauthors=Šarac J, Šarić T, Havaš Auguštin D, Novokmet N, Vekarić N, Mustać M, Grahovac B, Kapović M, Nevajda B, Glasnović A, Missoni S, Rootsi S, Rudan P | title=Genetic heritage of Croatians in the Southeastern European gene pool-Y chromosome analysis of the Croatian continental and Island population | journal=American Journal of Human Biology | volume=28 | issue=6 | pages=837–845 | year=2016 | pmid=27279290 | doi=10.1002/ajhb.22876 | s2cid=25873634 }} | ||||||||||||||||||||||||||||||||||||
| Northern Portugueses | Portuguese (Romance) | Vila Real | 3/39 | 7.7% | vauthors=Beleza S, Gusmão L, Lopes A, Alves C, Gomes I, Giouzeli M, Calafell F, Carracedo A, Amorim A | title=Micro-phylogeographic and demographic history of Portuguese male lineages | journal=Annals of Human Genetics | volume=70 | issue=Pt 2 | pages=181–94 | year=2006 | pmid=16626329 | doi=10.1111/j.1529-8817.2005.00221.x | s2cid=4652154 }} | ||||||||||||||||||||||||||||||||||||
| Materanis | Neapolitan language (Romance) | Matera and Policoro | 4/52 | 7.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Campanians | Neapolitan language (Romance) | Campania | 8/108 | 7.4% | vauthors=Calcagno G, Labruna G, Sacchetti L | title=Y-chromosome short tandem repeat (STR) haplotypes in a Campania population sample | journal=Clinical Chemistry and Laboratory Medicine | volume=43 | issue=2 | pages=163–6 | year=2005 | pmid=15843210 | doi=10.1515/CCLM.2005.027 | s2cid=43323602 }} | ||||||||||||||||||||||||||||||||||||
| Cretans | Cretan Greek | Oropedio Lasithiou | 3/41 | 7.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Latinensis | Neapolitan language (Romance) (Romance) | Norma and Sezze | 3/41 | 7.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Ragusa | 2/28 | 7.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Piazza Armerina | 2/28 | 7.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Trapani | 3/43 | 7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ligurians | Ligurian (Romance) | La Spezia | 3/43 | 7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Leccesis | Salentino language (Romance) | Vaste and Ugento | 3/46 | 6.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Walloons | Walloon (Romance) | Wallonia | 3/47 | 6.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ascolanis | Marchigiano (Romance) | Offida and Ascoli Piceno | 3/47 | 6.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Asturianos | Eonavian (Romance) | Navia-Eo | 2/31 | 6.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gagauzes | Gagauz (Turkic) | Kongaz | 3/48 | 6.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Solàndris | Solànder (Rhaeto-Romance) | Val de Sól | 4/65 | 6.2% | vauthors=Coia V, Capocasa M, Anagnostou P, Pascali V, Scarnicci F, Boschi I, Battaggia C, Crivellaro F, Ferri G, Alù M, Brisighelli F, Busby GB, Capelli C, Maixner F, Cipollini G, Viazzo PP, Zink A, Destro Bisol G | title=Demographic histories, isolation and social factors as determinants of the genetic structure of Alpine linguistic groups | journal=PLOS ONE | volume=8 | issue=12 | article-number=e81704 | year=2013 | pmid=24312576 | pmc=3847036 | doi=10.1371/journal.pone.0081704 | bibcode=2013PLoSO...881704C | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Northern Portuguese | Portuguese (Romance) | Aveiro | 4/66 | 6.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Western Andalusians | Andalusian (Romance) | Huelva | 10/167 | 6% | vauthors=Ambrosio B, Novelletto A, Hernandez C, Dugoujon JM, Fortes-Lima C, Rodriguez JN, Calderon R | title=Y-STR genetic diversity in autochthonous Andalusians from Huelva and Granada provinces (Spain) | journal=Forensic Science International. Genetics | volume=6 | issue=2 | pages=e66–71 | year=2012 | pmid=21664894 | doi=10.1016/j.fsigen.2011.05.007}} | |||||||||||||||||||||||||||||||||||||
| Aragonese | Aragonese and Castilian (Romance) | Aragón | 2/34 | 5.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Corsicans | Corsican | Corsica | 2/34 | 5.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Panteschis | Sicilian with Siculo-Arabic influences (Romance) | Pantelleria | 1/17 | 5.9% | vauthors=Robino C, Gino S, Ricci U, Grignani P, Previdere C, Torre C | title=Y-chromosomal STR haplotypes in an Albanian population sample | journal=Forensic Science International | volume=129 | issue=2 | pages=128–30 | year=2002 | pmid=12243882 | doi=10.1016/S0379-0738(02)00224-4}} | |||||||||||||||||||||||||||||||||||||
| Extremadurans | Astur-Leonese and Castilian (Romance) | Extremadura | 3/52 | 5.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Unspecified Bulgarian region | 4/69 | 5.8% | vauthors=Karachanak S, Grugni V, Fornarino S, Nesheva D, Al-Zahery N, Battaglia V, Carossa V, Yordanov Y, Torroni A, Galabov AS, Toncheva D, Semino O | title=Y-chromosome diversity in modern Bulgarians: new clues about their ancestry | journal=PLOS ONE | volume=8 | issue=3 | article-number=e56779 | year=2013 | pmid=23483890 | pmc=3590186 | doi=10.1371/journal.pone.0056779 | bibcode=2013PLoSO...856779K | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Tuscans | Tuscan (Romance) | Tuscany | 3/53 | 5.7% | vauthors=Poznik GD, Xue Y, Mendez FL, Willems TF, Massaia A, Wilson Sayres MA, Ayub Q, McCarthy SA, Narechania A, Kashin S, Chen Y, Banerjee R, Rodriguez-Flores JL, Cerezo M, Shao H, Gymrek M, Malhotra A, Louzada S, Desalle R, Ritchie GR, Cerveira E, Fitzgerald TW, Garrison E, Marcketta A, Mittelman D, Romanovitch M, Zhang C, Zheng-Bradley X, Abecasis GR, McCarroll SA, Flicek P, Underhill PA, Coin L, Zerbino DR, Yang F, Lee C, Clarke L, Auton A, Erlich Y, Handsaker RE, Bustamante CD, Tyler-Smith C | title=Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences | journal=Nature Genetics | volume=48 | issue=6 | pages=593–9 | year=2016 | pmid=27111036 | pmc=4884158 | doi=10.1038/ng.3559 | bibcode=2016NaGen..48..593P }} | |||||||||||||||||||||||||||||||||||
| Dutch | Hollandic (West Germanic) | North Holland | 1/18 | 5.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lombardians | Lombard and Italian (Romance) | Lombardia | 1/18 | 5.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | Mazara del Vallo | 1/18 | 5.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Southern Italians | Italian (Romance) | South Apulia | 4/71 | 5.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Asturians | Astur-Leonese (Romance) | Asturies | 4/74 | 5.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | South Sicily | 3/55 | 5.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lombardians | Lombard and Italian (Romance) | Lombardia | 7/131 | 5.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Hutterites | Austro-Bavarian (Upper German) | South Tyrol | 4/75 | 5.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Peloponnesians | Southern Greek | Peloponnese | 1/19 | 5.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gutes | Gutnish (North Germanic) | Gotland | 2/40 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Alsatians | Alsatian (Upper German) | Strossburi | 4/80 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Asturians | Astur-Leonese (Romance) | Asturies | 1/20 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Italian speakers | Italian (Romance) | Bozen | 3/59 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ladin Stilfser/Tyrolese | Ladin (Romance) | Stelvio | 1/20 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gaditanos | Andalusian language (Romance) | Cádiz | 1/20 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Malacitanos | Andalusian language (Romance) | Málaga | 1/20 | 5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Macedonians and Thracians | Northern Greek | East Macedonia and Thrace | 1/21 | 4.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Razgrad | 1/21 | 4.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northeastern Portuguese | Portuguese (Romance) | Trás os Montes | 3/64 | 4.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Corsicans | Gallurese (Romance languages) | Tempiu | 4/86 | 4.7% | vauthors=Contu D, Morelli L, Santoni F, Foster JW, Francalacci P, Cucca F | title=Y-chromosome based evidence for pre-neolithic origin of the genetically homogeneous but diverse Sardinian population: inference for association scans | journal=PLOS ONE | volume=3 | issue=1 | article-number=e1430 | year=2008 | pmid=18183308 | pmc=2174525 | doi=10.1371/journal.pone.0001430 | bibcode=2008PLoSO...3.1430C | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Sardinians | Sassarese (Romance) | Sassari | 2/43 | 4.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Jennesis | Central Italian language (Romance) | Jenne | 3/65 | 4.6% | Isolated mountain community | |||||||||||||||||||||||||||||||||||||||||||||
| Aretuseis | Sicilian (Romance) | Buccheri | 1/22 | 4.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Casteddammaresis | Sicilian (Romance) | Casteddammari | 1/22 | 4.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | East Sicily | 4/87 | 4.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Western Andalusians | Andalusian (Romance) | Huelva | 1/22 | 4.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| West Andalusians | Andalusian (Romance) | Sevilla | 7/155 | 4.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Galicians | Galician (Romance) | Santiago | 2/46 | 4.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Palentinos | Castilian language (Romance) | Palencia | 1/23 | 4.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Aragó | 1/23 | 4.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ligurians | Ligurian (Romance) | Central Liguria | 2/45 | 4.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Penedès | 7/164 | 4.3% | vauthors=Solé-Morata N, Bertranpetit J, Comas D, Calafell F | title=Y-chromosome diversity in Catalan surname samples: insights into surname origin and frequency | journal=European Journal of Human Genetics | volume=23 | issue=11 | pages=1549–57 | year=2015 | pmid=25689924 | pmc=4613475 | doi=10.1038/ejhg.2015.14}} | ||||||||||||||||||||||||||||||||||||
| Greeks | Greek | Athens | 4/92 | 4.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northern Portuguese | Portuguese | Beira Litoral | 5/116 | 4.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ligurians | Ligurian (Romance) | La Spezia | 2/46 | 4.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| South Italians | Salentino (Romance) | North Apulia | 2/46 | 4.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cantabrians | Astur-Leonese (Romance) | Cantabria | 3/70 | 4.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cimbrians | Cimbrian (West Germanic languages) | Lessinia | 1/24 | 4.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Pincianos | Castilian language (Romance) | Valladolid | 1/24 | 4.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Croats | Croatian (West Slavic) | Zadar Hinterland | 1/25 | 4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Macedonians | Northern Greek | Central Macedonia | 1/25 | 4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Madrileños | Castilian language (Romance) | Madrid | 2/50 | 4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Germans | German (West Germanic) | Berlin | 4/103 | 3.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northern Portuguese | Portuguese (Romance) | Braga | 2/51 | 3.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Beneventanis | Neapolitan language (Romance) | San Giorgio la Molara | 1/26 | 3.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Tuscans | Tuscan (Romance) | South Tuscany | 3/79 | 3.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Riojans | Riojan and Castilian (Romance) | La Rioja | 2/54 | 3.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Marchigianos | Marchigiano (Romance) | Apennines Marche | 1/27 | 3.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Calabrians | Southern Italian (Romance) | West Calabria | 1/27 | 3.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Urban Biellesi | Piedmontese (Romance) | Bièla | 3/81 | 3.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ukrainians | Ukrainian (East Slavic) | Kharkiv Oblast | 2/55 | 3.6% | vauthors=Kushniarevich A, Utevska O, Chuhryaeva M, Agdzhoyan A, Dibirova K, Uktveryte I, Möls M, Mulahasanovic L, Pshenichnov A, Frolova S, Shanko A, Metspalu E, Reidla M, Tambets K, Tamm E, Koshel S, Zaporozhchenko V, Atramentova L, Kučinskas V, Davydenko O, Goncharova O, Evseeva I, Churnosov M, Pocheshchova E, Yunusbayev B, Khusnutdinova E, Marjanović D, Rudan P, Rootsi S, Yankovsky N, Endicott P, Kassian A, Dybo A, Tyler-Smith C, Balanovska E, Metspalu M, Kivisild T, Villems R, Balanovsky O | title=Genetic Heritage of the Balto-Slavic Speaking Populations: A Synthesis of Autosomal, Mitochondrial and Y-Chromosomal Data | journal=PLOS ONE | volume=10 | issue=9 | article-number=e0135820 | year=2015 | pmid=26332464 | pmc=4558026 | doi=10.1371/journal.pone.0135820 | bibcode=2015PLoSO..1035820K | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Native Sayaguese speakers | Astur-Leonese (Romance) | Sayago | 1/28 | 3.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Galicians | Galician (Romance) | Montes Baixo Miño | 1/28 | 3.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Corsicans | Corsican (Romance) | Ajaccio (region of Corsica sutana) | 1/28 | 3.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sardinians | Sardinian (Romance) | Sassari and Orgosolo | 2/56 | 3.6% | vauthors=Verzeletti A, Cerri N, Gasparini F, Poglio A, Mazzeo E, De Ferrari F | title=Population data for 15 autosomal STRs loci and 12 Y chromosome STRs loci in a population sample from the Sardinia island (Italy) | journal=Legal Medicine | volume=11 | issue=1 | pages=37–40 | year=2009 | pmid=18723383 | doi=10.1016/j.legalmed.2008.06.003}} | |||||||||||||||||||||||||||||||||||||
| Southern Portugueses | Portuguese (Romance) | Évora | 1/29 | 3.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cretans | Cretan Greek | Khania | 1/29 | 3.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Canarians | Canarian Spanish (Romance) | La Palma | 3/85 | 3.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Scanians | Scanian dialects (South Scandinavian) | Malmö | 1/29 | 3.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Auvergnats | Auvergnat (Romance) | Clermont-Ferrand | 3/89 | 3.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Azoreans | Portuguese (Romance) | Eastern Azores | 3/87 | 3.4% | vauthors=Montiel R, Bettencourt C, Silva C, Santos C, Prata MJ, Lima M | title=Analysis of Y-chromosome variability and its comparison with mtDNA variability reveals different demographic histories between islands in the Azores Archipelago (Portugal) | journal=Annals of Human Genetics | volume=69 | issue=Pt 2 | pages=135–44 | year=2005 | pmid=15720295 | doi=10.1046/j.1529-8817.2004.00146.x | doi-broken-date=7 September 2025 | hdl=10316/8064 | s2cid=26096151 | hdl-access=free }} | |||||||||||||||||||||||||||||||||
| Asturians | Astur-Leonese (Romance) | Uviéu | 6/182 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Galicians | Galician (Romance) | Lugo | 2/61 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Albanians | Albanian dialects | Albania | 1/30 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northeastern Portuguese | Portuguese (Romance) | Bragança | 1/30 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northern Portuguese | Portuguese (Romance) | Viseu | 1/30 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Northern Portuguese | Portuguese (Romance) | Guarda | 1/30 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catanzaresis | southern Calabrese (Romance) | Catanzaro | 1/30 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sicilians | Sicilian (Romance) | West Sicily | 4/122 | 3.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Leoneses | Astur-leonese language (Romance) | Leon | 7/221 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lithuanians | Aukštaitian (Baltic) | West Aukstaiciai | 1/31 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Euboeans | Thessalian (Hellenic) | Euboea | 3/93 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Greeks | Northern Greek | Western Greece | 1/31 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Campanians | Neapolitan language (Romance) | San Giorgio La Molara | 1/31 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Valencians | Catalan and Castilian (Romance) | Valencia | 1/31 | 3.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Southern Tyroleans | Southern Austro-Bavarian (Upper German) | Lower Vinschgau | 1/32 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Rhinelanders | Ripuarian (Central Franconian) | Köln | 3/96 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Swedes | Swedish dialects (East Scandinavian) | Örebro | 1/32 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Cantabrians | Astur-Leonese (Romance) | Cantabria | 3/98 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Albaceteño | Castilian language (Romance) | Albacete | 1/32 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Portuguese | Portuguese (Romance) | Madeira | 4/129 | 3.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Asturianos | Astur-Leonese language (Romance) | Asturias | 1/33 | 3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lentinesi | Sicilian (Romance) | Lentini | 1/33 | 3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Shetlanders with Aboriginal surnames | Scots language and Norn Language (Germanic) | Shetland | 1/35 | 2.9% | Shetland Project | |||||||||||||||||||||||||||||||||||||||||||||
| Aretuseis | Sicilian (Romance) | Siracusa | 4/138 | 2.9% | last1=Tofanelli | first1=Sergio | last2=Brisighelli | first2=Francesca | last3=Anagnostou | first3=Paolo | last4=Busby | first4=George B. J. | last5=Ferri | first5=Gianmarco | last6=Thomas | first6=Mark G. | last7=Taglioli | first7=Luca | last8=Rudan | first8=Igor | last9=Zemunik | first9=Tatijana | last10=Hayward | first10=Caroline | last11=Bolnick | first11=Deborah | last12=Romano | first12=Valentino | last13=Cali | first13=Francesco | last14=Luiselli | first14=Donata | last15=Shepherd | first15=Gillian B. | last16=Tusa | first16=Sebastiano | last17=Facella | first17=Antonino | last18=Capelli | first18=Cristian | year=2015 | title=The Greeks in the West: genetic signatures of the Hellenic colonisation in southern Italy and Sicily | journal=European Journal of Human Genetics | volume=24 | issue=3 | pages=429–436 | doi=10.1038/ejhg.2015.124 | pmid=26173964 | pmc=4757772}} | |
| Baslers | Basel German (West Germanic) | Basel-Stadt | 18/643 | 2.8% | vauthors=Purps J, Siegert S, Willuweit S, Nagy M, Alves C, Salazar R, Angustia SM, Santos LH, Anslinger K, Bayer B, Ayub Q, Wei W, Xue Y, Tyler-Smith C, Bafalluy MB, Martínez-Jarreta B, Egyed B, Balitzki B, Tschumi S, Ballard D, Court DS, Barrantes X, Bäßler G, Wiest T, Berger B, Niederstätter H, Parson W, Davis C, Budowle B, Burri H, Borer U, Koller C, Carvalho EF, Domingues PM, Chamoun WT, Coble MD, Hill CR, Corach D, Caputo M, D'Amato ME, Davison S, Decorte R, Larmuseau MH, Ottoni C, Rickards O, Lu D, Jiang C, Dobosz T, Jonkisz A, Frank WE, Furac I, Gehrig C, Castella V, Grskovic B, Haas C, Wobst J, Hadzic G, Drobnic K, Honda K, Hou Y, Zhou D, Li Y, Hu S, Chen S, Immel UD, Lessig R, Jakovski Z, Ilievska T, Klann AE, García CC, de Knijff P, Kraaijenbrink T, Kondili A, Miniati P, Vouropoulou M, Kovacevic L, Marjanovic D, Lindner I, Mansour I, Al-Azem M, Andari AE, Marino M, Furfuro S, Locarno L, Martín P, Luque GM, Alonso A, Miranda LS, Moreira H, Mizuno N, Iwashima Y, Neto RS, Nogueira TL, Silva R, Nastainczyk-Wulf M, Edelmann J, Kohl M, Nie S, Wang X, Cheng B, Núñez C, Pancorbo MM, Olofsson JK, Morling N, Onofri V, Tagliabracci A, Pamjav H, Volgyi A, Barany G, Pawlowski R, Maciejewska A, Pelotti S, Pepinski W, Abreu-Glowacka M, Phillips C, Cárdenas J, Rey-Gonzalez D, Salas A, Brisighelli F, Capelli C, Toscanini U, Piccinini A, Piglionica M, Baldassarra SL, Ploski R, Konarzewska M, Jastrzebska E, Robino C, Sajantila A, Palo JU, Guevara E, Salvador J, Ungria MC, Rodriguez JJ, Schmidt U, Schlauderer N, Saukko P, Schneider PM, Sirker M, Shin KJ, Oh YN, Skitsa I, Ampati A, Smith TG, Calvit LS, Stenzl V, Capal T, Tillmar A, Nilsson H, Turrina S, De Leo D, Verzeletti A, Cortellini V, Wetton JH, Gwynne GM, Jobling MA, Whittle MR, Sumita DR, Wolańska-Nowak P, Yong RY, Krawczak M, Nothnagel M, Roewer L | title=A global analysis of Y-chromosomal haplotype diversity for 23 STR loci | journal=Forensic Science International. Genetics | volume=12 | pages=12–23 | year=2014 | issue=100 | pmid=24854874 | pmc=4127773 | doi=10.1016/j.fsigen.2014.04.008}} | ||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Smolensk Oblast | 3/107 | 2.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gienenses | Castilian language (Romance) | Jaen | 1/36 | 2.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Native Alistano speakers | Astur-Leonese (Romance) | Aliste | 1/36 | 2.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Germans | German (Germanic) | Germany | 1/37 | 2.7% | Karafet15 | |||||||||||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Oryol Oblast | 3/110 | 2.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Macedonians | Macedonian (Balto-Slavic) | Macedonia | 4/150 | 2.7% | vauthors=Spiroski M, Arsov T, Krüger C, Willuweit S, Roewer L | title=Y-chromosomal STR haplotypes in Macedonian population samples | journal=Forensic Science International | volume=148 | issue=1 | pages=69–73 | year=2005 | pmid=15607593 | doi=10.1016/j.forsciint.2004.04.067}} | |||||||||||||||||||||||||||||||||||||
| Azoreans | Portuguese (Romance) | Central Azores | 2/76 | 2.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Augustanis | Sicilian (Romance) | Augusta | 1/38 | 2.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Czechs | Czech (West Slavic) | Vysočina Region | 1/40 | 2.5% | vauthors=Zastera J, Roewer L, Willuweit S, Sekerka P, Benesova L, Minarik M | title=Assembly of a large Y-STR haplotype database for the Czech population and investigation of its substructure | journal=Forensic Science International. Genetics | volume=4 | issue=3 | pages=e75–8 | year=2010 | pmid=20215022 | doi=10.1016/j.fsigen.2009.06.005}} | |||||||||||||||||||||||||||||||||||||
| Fiemmeses | Fiamazzo (Romance) | Val de Fiem | 1/41 | 2.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Flemish | Dutch (West Germanic) | Turnhout | 1/42 | 2.4% | vauthors=Larmuseau MH, Ottoni C, Raeymaekers JA, Vanderheyden N, Larmuseau HF, Decorte R | title=Temporal differentiation across a West-European Y-chromosomal cline: genealogy as a tool in human population genetics | journal=European Journal of Human Genetics | volume=20 | issue=4 | pages=434–40 | year=2012 | pmid=22126748 | pmc=3306861 | doi=10.1038/ejhg.2011.218}} | "1675" data set | |||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Oryol Oblast | 1/42 | 2.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Haskovo | 1/41 | 2.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Genoese Tabarkini | Ligurian (Romance languages) | U Pàize | 1/41 | 2.4% | vauthors=Robledo R, Corrias L, Bachis V, Puddu N, Mameli A, Vona G, Calò CM | title=Analysis of a genetic isolate: the case of Carloforte (Italy) | journal=Human Biology | volume=84 | issue=6 | pages=735–54 | year=2012 | pmid=23959646 | doi=10.3378/027.084.0602 | s2cid=6609913 | url=https://digitalcommons.wayne.edu/cgi/viewcontent.cgi?article=1027&context=humbiol_preprints | url-access=subscription }} | ||||||||||||||||||||||||||||||||||
| Genoese Tabarkini | Ligurian (Romance languages) | U Pàize | 1/48 | 2.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Flemish | Dutch (West Germanic) | Tongeren | 1/43 | 2.3% | author1-link=Maarten Larmuseau | vauthors=Larmuseau MH, Boon N, Vanderheyden N, Van Geystelen A, Larmuseau HF, Matthys K, De Clercq W, Decorte R | title=High Y-chromosomal diversity and low relatedness between paternal lineages on a communal scale in the Western European Low Countries during the surname establishment | journal=Heredity | volume=115 | issue=1 | pages=3–12 | year=2015 | pmid=25873146 | pmc=4815499 | doi=10.1038/hdy.2015.5 | bibcode=2015Hered.115....3L }} | T1a1a-L208 | |||||||||||||||||||||||||||||||||
| Sardinians | Sardinian, Corsican (Romance) | Sardinia | 28/1204 | 2.3% | vauthors=Francalacci P, Morelli L, Angius A, Berutti R, Reinier F, Atzeni R, Pilu R, Busonero F, Maschio A, Zara I, Sanna D, Useli A, Urru MF, Marcelli M, Cusano R, Oppo M, Zoledziewska M, Pitzalis M, Deidda F, Porcu E, Poddie F, Kang HM, Lyons R, Tarrier B, Gresham JB, Li B, Tofanelli S, Alonso S, Dei M, Lai S, Mulas A, Whalen MB, Uzzau S, Jones C, Schlessinger D, Abecasis GR, Sanna S, Sidore C, Cucca F | title=Low-pass DNA sequencing of 1200 Sardinians reconstructs European Y-chromosome phylogeny | journal=Science | volume=341 | issue=6145 | pages=565–9 | year=2013 | pmid=23908240 | doi=10.1126/science.1237947 | pmc=5500864 | bibcode=2013Sci...341..565F }} | |||||||||||||||||||||||||||||||||||
| Croats | Croatian (West Slavic) | Dubrovnik | 4/179 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Kursk Oblast | 1/45 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sardinians | Gallurese (Romance) | Gaddùra | 1/46 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sardinians | Sardinian (Romance) | Sardinia | 27/1204 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Belvederesi | Neapolitan language (Romance) | Belvedere Marittimo | 1/45 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Fascians | Fascian (Rhaeto-Romance) | Fascia | 1/47 | 2.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Lipetsk Oblast | 1/47 | 2.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ukrainians | Ukrainian (East Slavic) | Chernihiv Raion | 2/96 | 2.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sardinians | Campidanese (Romance) | Trexenta | 1/47 | 2.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Sardinians | Logudorese (Romance languages) | Benetuti | 1/48 | 2.1% | vauthors=Robledo R, Mameli A, Scudiero CM, Vona G, Corrias L, Bachis V, Culigioni C, Calò CM | title=Non-random distribution of 17 Y-chromosome STR loci in different areas of Sardinia | journal=Forensic Science International. Genetics | volume=16 | pages=26–8 | year=2015 | pmid=25498479 | doi=10.1016/j.fsigen.2014.11.019}} | ||||||||||||||||||||||||||||||||||||||
| Lithuanians | Aukštaitian (Baltic) | western Aukštaitija | 1/50 | 2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ukrainians | Ukrainian (East Slavic) | Sumy Oblast | 2/101 | 2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Zamoranos | Castilian (Romance) | Campos - Pan | 1/50 | 2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Southwestern Almerians | Andalusian (Romance) | Laujar de Andarax, Ohanes, Berja and Adra | 1/50 | 2% | vauthors=Gaibar M, Esteban E, Moral P, Gómez-Gallego F, Santiago C, Bandrés F, Luna F, Fernández-Santander A | title=STR genetic diversity in a Mediterranean population from the south of the Iberian Peninsula | journal=Annals of Human Biology | volume=37 | issue=2 | pages=253–66 | year=2010 | pmid=19961347 | doi=10.3109/03014460903341851 | s2cid=19323185 }} | ||||||||||||||||||||||||||||||||||||
| Alpujarreños | Andalusian (Romance) | Alpujarra de la Sierra | 1/50 | 2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Corinthians | Ionian-Peloponesian and Albanian (Hellenic) | Corinthia | 2/104 | 1.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Macedonians | Macedonian (Balto-Slavic) | Macedonia | 4/211 | 1.9% | vauthors=Noveski P, Trivodalieva S, Efremov G, Plaseska-Karanfilska D | title=Y Chromosome Single Nucleotide Polymorphisms Typing by SNaPshot MINISEQUENCING | journal=Balkan Journal of Medical Genetics | volume=13 | issue=1 | pages=9–16 | year=2010 | doi=10.2478/v10034-010-0013-9 | doi-access=free }} | |||||||||||||||||||||||||||||||||||||
| Sardinians | Campidanese (Romance languages) | Sòrgono | 2/103 | 1.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan language (Romance language) | Camp de Tarragona | 4/214 | 1.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Ukrainians | Ukrainian (East Slavic) | Cherkasy Raion | 2/114 | 1.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Adigeses | Italian (Romance) | Val d'Adige | 1/56 | 1.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bosch surname members | Catalan language (Romance language) | Països Catalans | 1/56 | 1.8% | last1=Calafell | first1=Francesc | display-authors = etal | year=2013 | title=Estudi genètic dels cognoms catalans, valencians i balears | journal=Csic-Upf }} | ||||||||||||||||||||||||||||||||||||||||
| Basques | Gipuzkoan (Isolate language) | Southwestern Gipuzkoa | 1/57 | 1.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Basques | Gipuzkoan (Isolate language) | Gipuzkoa | 1/58 | 1.7% | last1=Young | first1=Kristin L. | last2=Sun | first2=Guangyun | last3=Deka | first3=Ranjan | last4=Crawford | first4=Michael H. | year=2011 | title=Paternal Genetic History of the Basque Population of Spain | journal=Human Biology | volume=83 | issue=4 | pages=455–475 | doi=10.3378/027.083.0402 | pmid=21846204 | hdl=1808/16387 | s2cid=3191418 | hdl-access=free }} | |||||||||||||||||||||||||||
| Flemish | Dutch (West Germanic) | North Brabant | 2/119 | 1.7% | "1775" data set | |||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Sofia | 1/59 | 1.7% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Lovech | 1/62 | 1.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Balearics | Majorcan (Romance) | Mallorca | 2/129 | 1.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Czechs | Czech (West Slavic) | Plzeň | 1/62 | 1.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Mecklenburgers | East Low Saxon (West Germanic) | Rostock | 3/200 | 1.5% | last1=Seiberling | first1=Susann | year=2005 | title=Allelverteilung Y-chromosomaler Short Tandem Repeats in Vorpommern | type=PhD Thesis | publisher=Greifswald Universitätsbibliothek | oclc=846027643}} | |||||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Belgorod Oblast | 2/143 | 1.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Castelló | 2/146 | 1.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Plovdiv | 2/159 | 1.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Montana, Bulgaria | 1/80 | 1.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Central Catalonia | 3/230 | 1.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Barcelona | 3/231 | 1.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Barcelona Periphery | 3/235 | 1.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Belarusians | Ukrainian (East Slavic) | Eastern Belarus | 1/86 | 1.2% | vauthors=Kushniarevich A, Sivitskaya L, Danilenko N, Novogrodskii T, Tsybovsky I, Kiseleva A, Kotova S, Chaubey G, Metspalu E, Sahakyan H, Bahmanimehr A, Reidla M, Rootsi S, Parik J, Reisberg T, Achilli A, Hooshiar Kashani B, Gandini F, Olivieri A, Behar DM, Torroni A, Davydenko O, Villems R | title=Uniparental genetic heritage of belarusians: encounter of rare middle eastern matrilineages with a central European mitochondrial DNA pool | journal=PLOS ONE | volume=8 | issue=6 | article-number=e66499 | year=2013 | pmid=23785503 | pmc=3681942 | doi=10.1371/journal.pone.0066499 | bibcode=2013PLoSO...866499K | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Czechs | Czech (West Slavic) | Ústí nad Labem | 1/86 | 1.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Russians | Russian (East Slavic) | Penza Oblast | 1/81 | 1.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Faroese | Faroese (Germanic) | Faroe Islands | 1/89 | 1.1% | vauthors=Jorgensen TH, Buttenschön HN, Wang AG, Als TD, Børglum AD, Ewald H | title=The origin of the isolated population of the Faroe Islands investigated using Y chromosomal markers | journal=Human Genetics | volume=115 | issue=1 | pages=19–28 | year=2004 | pmid=15083358 | doi=10.1007/s00439-004-1117-7 | s2cid=6040039 }} | Grandfathers originated from various Faroese islands. | |||||||||||||||||||||||||||||||||||
| Sardinians | Campidanese (Romance languages) | Casteddu | 2/187 | 1.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Eastern Andalusians | Andalusian (Romance) | Granada | 2/180 | 1.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Moravian Valachs | Romanian language (Romance languages) | Moravian Wallachia | 1/94 | 1.1% | vauthors=Ehler E, Vane D, Stenzl V, Vancata V | title=Y-chromosomal diversity of the Valachs from the Czech Republic: model for isolated population in Central Europe | journal=Croatian Medical Journal | volume=52 | issue=3 | pages=358–67 | year=2011 | pmid=21674832 | pmc=3131682 | doi=10.3325/cmj.2011.52.358}} | ||||||||||||||||||||||||||||||||||||
| Belarusians | Ukrainian (East Slavic) | Eastern Polesie | 1/96 | 1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Estonians | Estonian (Uralic) | Estonia | 2/209 | 1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Austrians | Southern Bavarian (Germanic) | Salzburg (state) | 2/200 | 1% | vauthors=Pickrahn I, Müller E, Zahrer W, Dunkelmann B, Cemper-Kiesslich J, Kreindl G, Neuhuber F | title=Yfiler Plus amplification kit validation and calculation of forensic parameters for two Austrian populations | journal=Forensic Science International. Genetics | volume=21 | pages=90–4 | year=2016 | pmid=26741856 | doi=10.1016/j.fsigen.2015.12.014}} | ||||||||||||||||||||||||||||||||||||||
| Ukrainians | Ukrainian (East Slavic) | Lviv Oblast | 1/101 | 1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Aragonese | Aragonese and Castilian (Romance) | Aragón | 2/200 | 1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Castellonenses | Catalan language (Romance) | Castelló | 5/515 | 1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bavarians | Bavarian (Germanic) | Bavaria | 2/218 | 0.9% | vauthors=Rębała K, Martínez-Cruz B, Tönjes A, Kovacs P, Stumvoll M, Lindner I, Büttner A, Wichmann HE, Siváková D, Soták M, Quintana-Murci L, Szczerkowska Z, Comas D | title=Contemporary paternal genetic landscape of Polish and German populations: from early medieval Slavic expansion to post-World War II resettlements | journal=European Journal of Human Genetics | volume=21 | issue=4 | pages=415–22 | year=2013 | pmid=22968131 | pmc=3598329 | doi=10.1038/ejhg.2012.190}} | T1a1a1a1b1-PF7445 | |||||||||||||||||||||||||||||||||||
| Austrian Germans | Southern Bavarian (Germanic) | Upper Austria | 2/225 | 0.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Czechs | Czech (West Slavic) | South Moravia | 2/216 | 0.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Croatians | Croatian (West Slavic) | Zagreb | 1/114 | 0.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Girona | 2/219 | 0.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Belarusians | Ukrainian (East Slavic) | Western Polesie | 1/121 | 0.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Mecklenburger | Mecklenburgisch-Vorpommersch (Germanic) | Mecklenburg | 1/138 | 0.8% | T1a2b-L446(xCTS11984) DYS437=15 | |||||||||||||||||||||||||||||||||||||||||||||
| Bulgarians | Bulgarian language (South Slavic languages) | Sofia Province | 2/257 | 0.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Andalusians | Andalusian (Romance) | Huelva Seville Córdoba Jaén Málaga Cádiz Granada Almeria | 1/144 | 0.7% | vauthors=Rey-González D | title=Micro and macro geographical analysis of Y-chromosome lineages in South Iberia | journal=Forensic Science International. Genetics | volume=29 | pages=e9–e15 | year=2017 | pmid= 28487219 | doi=10.1016/j.fsigen.2017.04.021}} | ||||||||||||||||||||||||||||||||||||||
| Romanians | Romanian (Romance) | Romania | 1/178 | 0.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Catalans | Catalan (Romance) | Valencia | 1/173 | 0.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Slovaks | Slovak (West Slavic) | Slovakia | 1/164 | 0.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Irish | Gaeilge (Celtic) | Ireland | 1/221 | 0.5% | vauthors=Hill EW, etal | title=Y-chromosome variation and Irish origins | journal=Nature | volume= 404 | issue= 6776 | pages= 351–2 | year=2000 | pmid= 10746711 | doi=10.1038/35006158 | bibcode=2000Natur.404..351H | s2cid=4414538 }} | |||||||||||||||||||||||||||||||||||
| Czechs | Czech (West Slavic) | Prague | 3/595 | 0.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Germans | German (West Germanic) | area of Halle | 1/234 | 0.4% | vauthors=Immel UD, Kleiber M, Klintschar M | title=Y chromosome polymorphisms and haplotypes in South Saxony-Anhalt (Germany) | journal=Forensic Science International | volume=155 | issue=2–3 | pages=211–5 | year=2005 | pmid=16226160 | doi=10.1016/j.forsciint.2005.01.004}} | |||||||||||||||||||||||||||||||||||||
| Individuals living in Catalonia | Catalan language (Romance) | Barcelona metropolitan area | 1/247 | 0.4% | vauthors=Sánchez C, Barrot C, Xifró A, Ortega M, de Aranda IG, Huguet E, Corbella J, Gené M | title=Haplotype frequencies of 16 Y-chromosome STR loci in the Barcelona metropolitan area population using Y-Filer kit | journal=Forensic Science International | volume=172 | issue=2–3 | pages=211–7 | year=2007 | pmid=17320328 | doi=10.1016/j.forsciint.2007.01.007}} | |||||||||||||||||||||||||||||||||||||
| Slovaks | Slovak (West Slavic) | Slovakia | 1/473 | 0.2% |
With K-M9+, unconfirmed but probable T-M70+: 14% (3/23) of Russians in Yaroslavl, 12.5% (3/24) of Italians in Matera, 10.3% (3/29) of Italians in Avezzano, 10% (3/30) of Tyroleans in Nonstal, 10% (2/20) of Italians in Pescara, 8.7% (4/46) of Italians in Benevento, 7.8% (4/51) of Italians in South Latium, 7.4% (2/27) of Italians in Paola, 7.3% (11/150) of Italians in Central-South Italy, 7.1% (8/113) of Serbs in Serbia, 4.7% (2/42) of Aromanians in Romania, 3.7% (3/82) of Italians in Biella, 3.7% (1/27) of Andalusians in Córdoba, 3.3% (2/60) of Leoneses in León, 3.2% (1/31) of Italians in Postua, 3.2% (1/31) of Italians in Cavaglià, 3.1% (3/97) of Calabrians in Reggio Calabria, 2.8% (1/36) of Russians in Ryazan Oblast, 2.8% (2/72) of Italians in South Apulia, 2.7% (1/37) of Calabrians in Cosenza, 2.6% (3/114) of Serbs in Belgrade, 2.5% (1/40) of Russians in Pskov, 2.4% (1/42) of Russians in Kaluga, 2.2% (2/89) of Transylvanians in Miercurea Ciuc, 2.2% (2/92) of Italians in Trino Vercellese, 1.9% (2/104) of Italians in Brescia, 1.9% (2/104) of Romanians in Romania, 1.7% (4/237) of Serbs and Montenegrins in Serbia and Montenegro, 1.7% (1/59) of Italians in Marche, 1.7% (1/59) of Calabrians in Catanzaro, 1.6% (3/183) of Greeks in Northern Greece, 1.3% (2/150) of Swiss Germans in Zürich Area, 1.3% (1/79) of Italians in South Tuscany and North Latium, 1.1% (1/92) of Dutch in Leiden, 0.5% (1/185) of Serbs in Novi Sad (Vojvodina), 0.5% (1/186) of Polish in Podlasie
Other parts that have been found to contain a significant proportion of haplogroup T-M184 individuals include Trentino (2/67 or 3%), Mariña Lucense (1/34 or 2.9%), Heraklion (3/104 or 2.9%), Roslavl (3/107 or 2.8%), Ourense (1/37 or 2.7%), Livny (3/110 or 2.7%), Biella (3/114 or 2.6%), Entre Douro (6/228 or 2.6%), Porto (3/118 or 2.5%), Urbino (1/40 or 2.5%), Iberian Peninsula (16/629 or 2.5%), Blekinge/Kristianstad (1/41 or 2.4%), Belarus (1/41 or 2.4%), Modena (3/130 or 2.3%), Provence-Alpes-Côte d'Azur (1/45 or 2.2%), Pristen (1/45 or 2.2%), Cáceres (2/91 or 2.2%), Brac (1/47 or 2.1%), Satakunta (1/48 or 2.1%), Western Croatia (2/101 or 2%), Ukrainia (1/50 or 2%), Greifswald (2/104 or 1.9%), Moldavians in Sofia (1/54 or 1.9%), Uppsala (1/55 or 1.8%), Lublin (2/112 or 1.8%), Pias in Beja (1/54 or 1.8%), Macedonian Greeks (1/57 or 1.8%), Nea Nikomedeia (1/57 or 1.8%), Sesklo/Dimini (1/57 or 1.8%), Lerna/Franchthi (1/57 or 1.8%), Açores (2/121 or 1.7%), Viana do Castelo (1/59 or 1.7%), Toulouse (1/67 or 1.5%), Belgorod (2/143 or 1.4%), Sardinia (1/77 or 1.3%).
- According to data from commercial testing, 3.9% of Italian males belonging to this haplogroup. Approximately 3% of Sephardi Jews and 2% of Ashkenazi Jews belong to haplogroup T.
Middle East and Caucasus
Haplogroup T has some significant frequencies in southeast and eastern Anatolia, the Zagros Mountains and both sides of the Persian Gulf.
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | ||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Georgians | Georgian (Kartvelian) | Khashuri | 1/3 | 33.3% | ||||||||||||||||||||||||||||||||||||||||
| Priest Zoroastrians | Persian | Shiraz, Tehran and Yazd | 2/8 | 25% | vauthors=Lopez S, etal | title=The Genetic Legacy of Zoroastrianism in Iran and India: Insights into Population Structure, Gene Flow, and Selection | journal=The American Journal of Human Genetics | volume= 101 | issue= 3 | pages= 353–368 | year=2017 | pmid= 28844488 | doi=10.1016/j.ajhg.2017.07.013 | pmc=5590844}} | Not specified if Herbad or Mobad | |||||||||||||||||||||||||||||
| Iraqi Jews | Judeo-Iraqi Arabic (Central Semitic) | Iraq | 7/32 | 21.9% | 12.5% T1a1a1a1a1a1-P77 and 9.4% T1a3-Y11151 | |||||||||||||||||||||||||||||||||||||||
| Armenian Sasuntzis | Western Armenian dialect, Kurmanji and Dimli (Northwestern Iranian) languages | Sasun | 21/104 | 20.2% | vauthors=Herrera KJ, Lowery RK, Hadden L, Calderon S, Chiou C, Yepiskoposyan L, Regueiro M, Underhill PA, Herrera RJ | title=Neolithic patrilineal signals indicate that the Armenian plateau was repopulated by agriculturalists | journal=European Journal of Human Genetics | volume=20 | issue=3 | pages=313–20 | year=2012 | pmid=22085901 | pmc=3286660 | doi=10.1038/ejhg.2011.192}} | T1a1 and T1a2 subclades | |||||||||||||||||||||||||||||
| Georgians | Georgian (Kartvelian) | Sighnaghi and Gurjaani | 2/10 | 20% | ||||||||||||||||||||||||||||||||||||||||
| Georgians | Georgian (Kartvelian) | Kharagauli | 1/5 | 20% | vauthors=Tarkhnishvili D, Gavashelishvili A, Murtskhvaladze M, Gabelaia M, Tevzadze G | title=Human paternal lineages, languages, and environment in the Caucasus | journal=Human Biology | volume=86 | issue=2 | pages=113–30 | year=2014 | pmid=25397702 | doi=10.3378/027.086.0205 | s2cid=7733899 | url=https://digitalcommons.wayne.edu/cgi/viewcontent.cgi?article=1053&context=humbiol_preprints | url-access=subscription }} | ||||||||||||||||||||||||||||
| Kumyks | Kumyk (Turkic) | Daghestani lowlands | 2/10 | 20% | vauthors=Marchani EE, Watkins WS, Bulayeva K, Harpending HC, Jorde LB | title=Culture creates genetic structure in the Caucasus: autosomal, mitochondrial, and Y-chromosomal variation in Daghestan | journal=BMC Genetics | volume=9 | article-number=47 | year=2008 | pmid=18637195 | pmc=2488347 | doi=10.1186/1471-2156-9-47 | doi-access=free }} | Reported as K* but according to Karafet16 and Yunusbayev12 only T fits. | |||||||||||||||||||||||||||||
| Kurdish Jews | Judeo-Aramaic (Central Semitic) | Kurdistan | 19/99 | 19.2% | vauthors=Nebel A, Filon D, Brinkmann B, Majumder PP, Faerman M, Oppenheim A | title=The Y chromosome pool of Jews as part of the genetic landscape of the Middle East | journal=American Journal of Human Genetics | volume=69 | issue=5 | pages=1095–112 | year=2001 | pmid=11573163 | pmc=1274378 | doi=10.1086/324070}} | ||||||||||||||||||||||||||||||
| Kurdish Jews | Judeo-Aramaic (Central Semitic) | Kurdistan | 9/50 | 18% | 10% T1a1a1a1a1a1-P77 and 8% T1a1-L162 | |||||||||||||||||||||||||||||||||||||||
| Druzes | Palestinian Arabic (Central Semitic) | Galilee | 7/40 | 17.5% | ||||||||||||||||||||||||||||||||||||||||
| Assyrians | Aramaic (Central Semitic) | refugees in Armenia | 16/106 | 15.1% | vauthors=Yepiskoposian L, Khudoyan A, Harutyunian A | title=Genetic Testing of Language Replacement Hypothesis in Southwest Asia | journal=Iran and the Caucasus | volume=10 | issue=2 | year=2006 | pages=191–208 | jstor=4030922 | doi=10.1163/157338406780345899 | s2cid=162345193 }} | Reported as K*. Their homeland in the areas around Urmia. | |||||||||||||||||||||||||||||
| Assyrians | Aramaic (Central Semitic) | Unknown | 4/28 | 14.3% | ||||||||||||||||||||||||||||||||||||||||
| Georgians | Georgian (Kartvelian) | Dusheti | 1/7 | 14.3% | ||||||||||||||||||||||||||||||||||||||||
| Iranian Jews | Judeo-Iranian (Southwestern Iranian) | Iran | 3/22 | 13.6% | 4.5% T1a1a1a1a1a1-P77 and 9.1% T1a3-Y11151 | |||||||||||||||||||||||||||||||||||||||
| Zoroastrians | Persian | Kerman | 5/37 | 13.5% | vauthors=Lashgary Z, Khodadadi A, Singh Y, Houshmand SM, Mahjoubi F, Sharma P, Singh S, Seyedin M, Srivastava A, Ataee M, Mohammadi ZS, Rezaei N, Bamezai RN, Sanati MH | title=Y chromosome diversity among the Iranian religious groups: a reservoir of genetic variation | journal=Annals of Human Biology | volume=38 | issue=3 | pages=364–71 | year=2011 | pmid=21329477 | doi=10.3109/03014460.2010.535562 | s2cid=207460555 }} | ||||||||||||||||||||||||||||||
| Iraqi Jews | Judeo-Iraqi Arabic (Central Semitic) | Iraq | 13/99 | 13.1% | vauthors=Zoossmann-Diskin A | title=The origin of Eastern European Jews revealed by autosomal, sex chromosomal and mtDNA polymorphisms | journal=Biology Direct | volume=5 | page=57 | year=2010 | pmid=20925954 | pmc=2964539 | doi=10.1186/1745-6150-5-57 | doi-access=free }} | ||||||||||||||||||||||||||||||
| Bakhtiaris | Bakhtiari (Southwestern Iranian (Perside)) | Izeh | 13/103 | 12.6% | vauthors=Roewer L, Willuweit S, Stoneking M, Nasidze I | title=A Y-STR database of Iranian and Azerbaijanian minority populations | journal=Forensic Science International. Genetics | volume=4 | issue=1 | pages=e53–5 | year=2009 | pmid=19948326 | doi=10.1016/j.fsigen.2009.05.002}} | |||||||||||||||||||||||||||||||
| Mountain Jews | Judeo-Tat (Southwestern Iranian) | Derbentsky District | 2/17 | 11.8% | vauthors=Karafet TM, Bulayeva KB, Nichols J, Bulayev OA, Gurgenova F, Omarova J, Yepiskoposyan L, Savina OV, Rodrigue BH, Hammer MF | title=Coevolution of genes and languages and high levels of population structure among the highland populations of Daghestan | journal=Journal of Human Genetics | volume=61 | issue=3 | pages=181–91 | year=2016 | pmid=26607180 | doi=10.1038/jhg.2015.132 | s2cid=6641494 | url=http://www.escholarship.org/uc/item/5zr6g9fj | doi-access=free }} | All belong to T1a1a1a1a1a1-P77 | |||||||||||||||||||||||||||
| Armenians | Western Armenian dialect | Historical Southwestern Armenia | 11/96 | 11.5% | ||||||||||||||||||||||||||||||||||||||||
| Emiratis | Gulf Arabic (Semitic) | Abu Dhabi | 21/191 | 11% | W. Goodwin et al., " Department of Forensic and Investigative Science, " "http://www.yhrd.org/" (2012), | |||||||||||||||||||||||||||||||||||||||
| Assyrians | Assyrian (Central Semitic) | West Azerbaijan Province | 4/39 | 10.3% | vauthors=Grugni V, Battaglia V, Hooshiar Kashani B, Parolo S, Al-Zahery N, Achilli A, Olivieri A, Gandini F, Houshmand M, Sanati MH, Torroni A, Semino O | title=Ancient migratory events in the Middle East: new clues from the Y-chromosome variation of modern Iranians | journal=PLOS ONE | volume=7 | issue=7 | article-number=e41252 | year=2012 | pmid=22815981 | pmc=3399854 | doi=10.1371/journal.pone.0041252 | bibcode=2012PLoSO...741252G | doi-access=free }} | ||||||||||||||||||||||||||||
| Iranian Jews | Judeo-Iranian (Southwestern Iranian) | Iran | 5/49 | 10.2% | ||||||||||||||||||||||||||||||||||||||||
| Persian Muslims | Persian | Shiraz | 5/51 | 9.8% | ||||||||||||||||||||||||||||||||||||||||
| Persian Muslims | Persian | Kerman | 6/66 | 9.1% | ||||||||||||||||||||||||||||||||||||||||
| Iraqis | Iraqi Arabic (Semitic) | Al-Qadisiyah | 6/69 | 8.7% | ||||||||||||||||||||||||||||||||||||||||
| Armenians | Armenian | Armenia | 35/413 | 8.5% | doi=10.1038/srep20768 | |||||||||||||||||||||||||||||||||||||||
| Kurds | Sorani (Northwestern Iranian) | Kurdestan | 5/59 | 8.5% | ||||||||||||||||||||||||||||||||||||||||
| Omani Arabs | Omani Arabic (Semitic) | Oman | 10/121 | 8.3% | vauthors=Luis JR, Rowold DJ, Regueiro M, Caeiro B, Cinnioğlu C, Roseman C, Underhill PA, Cavalli-Sforza LL, Herrera RJ | title=The Levant versus the Horn of Africa: evidence for bidirectional corridors of human migrations | journal=American Journal of Human Genetics | volume=74 | issue=3 | pages=532–44 | year=2004 | pmid=14973781 | pmc=1182266 | doi=10.1086/382286}} | ||||||||||||||||||||||||||||||
| Kurds | Sorani (Northwestern Iranian) | Kurdestan | 2/25 | 8% | vauthors=Di Cristofaro J, Pennarun E, Mazières S, Myres NM, Lin AA, Temori SA, Metspalu M, Metspalu E, Witzel M, King RJ, Underhill PA, Villems R, Chiaroni J | title=Afghan Hindu Kush: where Eurasian sub-continent gene flows converge | journal=PLOS ONE | volume=8 | issue=10 | article-number=e76748 | year=2013 | pmid=24204668 | pmc=3799995 | doi=10.1371/journal.pone.0076748 | bibcode=2013PLoSO...876748D | doi-access=free }} | ||||||||||||||||||||||||||||
| Azeris | Azeri (Oghuz) | West Azerbaijan Province | 5/63 | 7.9% | ||||||||||||||||||||||||||||||||||||||||
| Mazanderanis | Mazanderan (Western Iranian) | Mazandaran | 1/13 | 7.7% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Cyprus | 3/41 | 7.3% | vauthors=Badro DA, Douaihy B, Haber M, Youhanna SC, Salloum A, Ghassibe-Sabbagh M, Johnsrud B, Khazen G, Matisoo-Smith E, Soria-Hernanz DF, Wells RS, Tyler-Smith C, Platt DE, Zalloua PA | title=Y-chromosome and mtDNA genetics reveal significant contrasts in affinities of modern Middle Eastern populations with European and African populations | journal=PLOS ONE | volume=8 | issue=1 | article-number=e54616 | year=2013 | pmid=23382925 | pmc=3559847 | doi=10.1371/journal.pone.0054616 | bibcode=2013PLoSO...854616B | doi-access=free }} | ||||||||||||||||||||||||||||
| Iraqis | Iraqi Arabic (Semitic) | Iraq | 10/139 | 7.2% | vauthors=Al-Zahery N, Semino O, Benuzzi G, Magri C, Passarino G, Torroni A, Santachiara-Benerecetti AS | title=Y-chromosome and mtDNA polymorphisms in Iraq, a crossroad of the early human dispersal and of post-Neolithic migrations | journal=Molecular Phylogenetics and Evolution | volume=28 | issue=3 | pages=458–72 | year=2003 | pmid=12927131 | doi=10.1016/s1055-7903(03)00039-3 | bibcode=2003MolPE..28..458A }} | ||||||||||||||||||||||||||||||
| Kuwaitis | Gulf Arabic (Semitic) | Kuwait | 3/42 | 7.1% | vauthors=El-Sibai M, Platt DE, Haber M, Xue Y, Youhanna SC, Wells RS, Izaabel H, Sanyoura MF, Harmanani H, Bonab MA, Behbehani J, Hashwa F, Tyler-Smith C, Zalloua PA | title=Geographical structure of the Y-chromosomal genetic landscape of the Levant: a coastal-inland contrast | journal=Annals of Human Genetics | volume=73 | issue=Pt 6 | pages=568–81 | year=2009 | pmid=19686289 | pmc=3312577 | doi=10.1111/j.1469-1809.2009.00538.x}} | ||||||||||||||||||||||||||||||
| Iraqis | Iraqi Arabic (Semitic) | Iraq | 3/43 | 7% | vauthors=Quintana-Murci L, Semino O, Poloni ES, Liu A, Van Gijn M, Passarino G, Brega A, Nasidze IS, Maccioni L, Cossu G, al-Zahery N, Kidd JR, Kidd KK, Santachiara-Benerecetti AS | title=Y-chromosome specific YCAII, DYS19 and YAP polymorphisms in human populations: a comparative study | journal=Annals of Human Genetics | volume=63 | issue=Pt 2 | pages=153–66 | year=1999 | pmid=10738527 | doi=10.1046/j.1469-1809.1999.6320153.x | s2cid=19675208 | doi-access=free }} | |||||||||||||||||||||||||||||
| Arabs | Levantine Arabic | Israel and Palestine | 10/143 | 7% | vauthors=Mukherjee N, Nebel A, Oppenheim A, Majumder PP | title=High-resolution analysis of Y-chromosomal polymorphisms reveals signatures of population movements from Central Asia and West Asia into India | journal=Journal of Genetics | volume=80 | issue=3 | pages=125–35 | year=2001 | pmid=11988631 | doi=10.1007/bf02717908 | s2cid=13267463 }} | ||||||||||||||||||||||||||||||
| Persians | Farsi (Southwestern Iranian) | Fars | 3/44 | 6.8% | ||||||||||||||||||||||||||||||||||||||||
| Christian Arabs | Levantine Arabic | Israel and Palestine | 3/44 | 6.8% | vauthors=Fernandes AT, Gonçalves R, Gomes S, Filon D, Nebel A, Faerman M, Brehm A | title=Y-chromosomal STRs in two populations from Israel and the Palestinian Authority Area: Christian and Muslim Arabs | journal=Forensic Science International. Genetics | volume=5 | issue=5 | pages=561–2 | year=2011 | pmid=20843760 | doi=10.1016/j.fsigen.2010.08.005 | hdl=10400.13/4485 | hdl-access=free }} | |||||||||||||||||||||||||||||
| Western Armenians | Armenian | Eastern Turkey | 6/90 | 6.7% | ||||||||||||||||||||||||||||||||||||||||
| Persians | Farsi (Southwestern Iranian) | Yazd | 3/46 | 6.5% | ||||||||||||||||||||||||||||||||||||||||
| Armenians | Armenian | Gardman | 6/96 | 6.3% | ||||||||||||||||||||||||||||||||||||||||
| Yezidis | Kurmanji (Northwestern Iranian) | refugees in Armenia | 12/196 | 6.1% | Reported as K*. Their homeland in the areas around Laliş. | |||||||||||||||||||||||||||||||||||||||
| Muslim Arabs | Levantine Arabic | Israel and Palestine | 7/119 | 5.9% | ||||||||||||||||||||||||||||||||||||||||
| Zahedan, Baluchestan, Iran | 6/103 | 5.8% | ||||||||||||||||||||||||||||||||||||||||||
| Northern Armenians | Armenian | Northern Armenia, southern Georgia (Bolnisi, Akhalkalaki and Akhaltsikhe) and northwestern Azerbaijan (around Gyanja) | 10/189 | 5.3% | ||||||||||||||||||||||||||||||||||||||||
| Armenians | Armenian | Tehran | 2/38 | 5.3% | ||||||||||||||||||||||||||||||||||||||||
| Eastern Armenians | Armenian | Karabakh | 11/215 | 5.1% | ||||||||||||||||||||||||||||||||||||||||
| Persians | Farsi (Southwestern Iranian) | Khorasan | 3/59 | 5.1% | ||||||||||||||||||||||||||||||||||||||||
| Saudi Arabians | Arabic dialects (Semitic) | Saudi Arabia | 8/157 | 5.1% | doi=10.1186/1471-2156-10-59 | title=Saudi Arabian Y-Chromosome diversity and its relationship with nearby regions | year=2009 | last1=Abu-Amero | first1=Khaled K | last2=Hellani | first2=Ali | last3=González | first3=Ana M | last4=Larruga | first4=Jose M | last5=Cabrera | first5=Vicente M | last6=Underhill | first6=Peter A | journal=BMC Genetics | volume=10 | article-number=59 | pmid=19772609 | pmc=2759955 | doi-access=free }} | |||||||||||||||||||
| Armenians | Armenian | Syunik | 7/140 | 5% | ||||||||||||||||||||||||||||||||||||||||
| Emiratis | Gulf Arabic (Semitic) | United Arab Emirates | 8/164 | 4.9% | ||||||||||||||||||||||||||||||||||||||||
| Lebanese Muslims | Lebanese Arabic (Semitic) | Lebanon | 28/568 | 4.9% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Lemesos | 6/126 | 4.8% | ||||||||||||||||||||||||||||||||||||||||
| Kumyks | Kumyk (Turkic) | Khasavyurtovsky District | 1/21 | 4.8% | ||||||||||||||||||||||||||||||||||||||||
| Avars | Avar (Northeast Caucasian) | southeastern Dagestan | 2/42 | 4.8% | ||||||||||||||||||||||||||||||||||||||||
| Kurds | Kurmanji (Northwestern Iranian) | Anatolia | 12/251 | 4.8% | vauthors=Flores C, Maca-Meyer N, Larruga JM, Cabrera VM, Karadsheh N, Gonzalez AM | title=Isolates in a corridor of migrations: a high-resolution analysis of Y-chromosome variation in Jordan | journal=Journal of Human Genetics | volume=50 | issue=9 | pages=435–41 | year=2005 | pmid=16142507 | doi=10.1007/s10038-005-0274-4 | doi-access=free }} | ||||||||||||||||||||||||||||||
| Kurds | Kurdish dialects (Northwestern Iranian) | Kurdistan | 6/126 | 4.8% | Carsten Hohoff and Bernd Brinkmann "Institut für Rechtsmedizin"," Universität Münster | |||||||||||||||||||||||||||||||||||||||
| Anizes | Gulf Arabic (Semitic) | Kuwait | 1/21 | 4.7% | vauthors=Mohammad T, Xue Y, Evison M, Tyler-Smith C | title=Genetic structure of nomadic Bedouin from Kuwait | journal=Heredity | volume=103 | issue=5 | pages=425–33 | year=2009 | pmid=19639002 | pmc=2869035 | doi=10.1038/hdy.2009.72 | bibcode=2009Hered.103..425M }} | |||||||||||||||||||||||||||||
| Lebaneses | Levantine Arabic (Semitic) | Lebanon | 43/914 | 4.7% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Cyprus | 3/65 | 4.6% | ||||||||||||||||||||||||||||||||||||||||
| Maronites | Lebanese Arabic and Syriac (Semitic) | Lebanon | 24/518 | 4.6% | ||||||||||||||||||||||||||||||||||||||||
| Armenians | Armenian | Ararat | 2/44 | 4.6% | vauthors=Weale ME, Yepiskoposyan L, Jager RF, Hovhannisyan N, Khudoyan A, Burbage-Hall O, Bradman N, Thomas MG | title=Armenian Y chromosome haplotypes reveal strong regional structure within a single ethno-national group | journal=Human Genetics | volume=109 | issue=6 | pages=659–74 | year=2001 | pmid=11810279 | doi=10.1007/s00439-001-0627-9 | s2cid=23113666 }} | ||||||||||||||||||||||||||||||
| Muslim Kurds | Kurdish dialects (Northwestern Iranian) | Kurdistan | 4/95 | 4.2% | ||||||||||||||||||||||||||||||||||||||||
| Qeshmis | Qishmi (southwestern Iranian) | Qeshm | 2/49 | 4.1% | ||||||||||||||||||||||||||||||||||||||||
| Lurs | Luri (Southwestern Iranian) | Lorestan | 2/50 | 4% | ||||||||||||||||||||||||||||||||||||||||
| Sadats | Languages of Iran | Different cities of Iran | 2/50 | 4% | vauthors=Rafiee MR, Sokhansanj A, Naghizadeh MA, Farazmand A | title=Analysis of Y-chromosomal short tandem repeat (STR) polymorphism in an Iranian Sadat population | journal=Russian Journal of Genetics | volume=45 | issue=8 | year=2009 | pages=969–73 | doi=10.1134/S1022795409080110 | pmid=19769300 | s2cid=24234321 }} | ||||||||||||||||||||||||||||||
| Persians | Persian | Eastern Iran | 3/77 | 3.9% | vauthors=Malyarchuk B, Derenko M, Wozniak M, Grzybowski T | title=Y-chromosome variation in Tajiks and Iranians | journal=Annals of Human Biology | volume=40 | issue=1 | pages=48–54 | year=2013 | pmid=23198991 | doi=10.3109/03014460.2012.747628 | s2cid=2752490 }} | ||||||||||||||||||||||||||||||
| Armenians | Armenian | Lake Van | 4/103 | 3.9% | ||||||||||||||||||||||||||||||||||||||||
| Saudi Arabians | Arabic dialects (Semitic) | Saudi Arabia | 4/106 | 3.8% | ||||||||||||||||||||||||||||||||||||||||
| Turkish Cypriots | Cypriot Turkish | 138 different villages, towns or cities from Cyprus | 14/380 | 3.7% | vauthors=Gurkan C, Sevay H, Demirdov DK, Hossoz S, Ceker D, Teralı K, Erol AS | title=Turkish Cypriot paternal lineages bear an autochthonous character and closest resemblance to those from neighbouring Near Eastern populations | journal=Annals of Human Biology | volume=44 | issue=2 | pages=164–174 | year=2017 | pmid=27356680 | doi=10.1080/03014460.2016.1207805 | s2cid=24596494 }} | Paternal lineages originating from the traditional Turkish Cypriot settlements throughout the island | |||||||||||||||||||||||||||||
| Birjand, South Khorasan, Iran | 1/27 | 3.7% | vauthors=Tabrizi AA, Hedjazi A, Kerachian MA, Honarvar Z, Dadgarmoghaddam M, Raoofian R | title=Genetic profile of 17 Y-chromosome STR haplotypes in East of Iran | journal=Forensic Science International. Genetics | volume=14 | pages=e6–7 | year=2015 | pmid=25458927 | doi=10.1016/j.fsigen.2014.10.010}} | All T1a3-Y12871 | |||||||||||||||||||||||||||||||||
| Armenians | Armenian | Ararat Valley | 4/110 | 3.6% | ||||||||||||||||||||||||||||||||||||||||
| Armenians | Armenian | Armenia | 2/57 | 3.5% | ||||||||||||||||||||||||||||||||||||||||
| Georgians | Georgian (Kartvelian) | Omalo | 1/29 | 3.5% | ||||||||||||||||||||||||||||||||||||||||
| Iranians | Languages of Iran | South Iran | 4/117 | 3.4% | doi=10.1159/000093774 | title=Iran: Tricontinental Nexus for Y-Chromosome Driven Migration | year=2006 | last1=Regueiro | first1=M. | last2=Cadenas | first2=A.M. | last3=Gayden | first3=T. | last4=Underhill | first4=P.A. | last5=Herrera | first5=R.J. | journal=Human Heredity | volume=61 | issue=3 | pages=132–43 | pmid=16770078 | s2cid=7017701 }} | |||||||||||||||||||||
| Ionians | Greek | Phokaia | 1/31 | 3.2% | doi=10.1186/1471-2148-11-69 | title=The coming of the Greeks to Provence and Corsica: Y-chromosome models of archaic Greek colonization of the western Mediterranean | year=2011 | last1=King | first1=Roy J | last2=Dicristofaro | first2=Julie | last3=Kouvatsi | first3=Anastasia | last4=Triantaphyllidis | first4=Costas | last5=Scheidel | first5=Walter | last6=Myres | first6=Natalie M | last7=Lin | first7=Alice A | last8=Eissautier | first8=Alexandre | last9=Mitchell | first9=Michael | last10=Binder | first10=Didier | last11=Semino | first11=Ornella | last12=Novelletto | first12=Andrea | last13=Underhill | first13=Peter A | last14=Chiaroni | first14=Jacques | journal=BMC Evolutionary Biology | volume=11 | issue=1 | page=69 | pmid=21401952 | pmc=3068964 | bibcode=2011BMCEE..11...69K | doi-access=free }} | |
| Bandaris | Bandari (Southwestern Iranian) | Bandar Abbas | 4/131 | 3.1% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Larnaka | 2/67 | 3% | ||||||||||||||||||||||||||||||||||||||||
| Alans | Karachay-Baksan-Chegem (Turkic) | Kabardino-Balkaria | 1/69 | 2.9% | ||||||||||||||||||||||||||||||||||||||||
| Jordanians | Arabic dialects (Semitic) | Jordan | 8/273 | 2.9% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Ammochostos | 3/122 | 2.5% | vauthors=Voskarides K, Mazières S, Hadjipanagi D, Di Cristofaro J, Ignatiou A, Stefanou C, King RJ, Underhill PA, Chiaroni J, Deltas C | title=Y-chromosome phylogeographic analysis of the Greek-Cypriot population reveals elements consistent with Neolithic and Bronze Age settlements | journal=Investigative Genetics | volume=7 | article-number=1 | year=2016 | pmid=26870315 | pmc=4750176 | doi=10.1186/s13323-016-0032-8 | doi-access=free }} | ||||||||||||||||||||||||||||||
| Lezghins | Lezgian (Northeast Caucasian) | Southern Dagestan | 2/81 | 2.5% | ||||||||||||||||||||||||||||||||||||||||
| Turks | Turkish | Turkey | 13/523 | 2.5% | ||||||||||||||||||||||||||||||||||||||||
| Persians | Persian (Southwestern Iranian) | Esfahan | 1/13 | 2.4% | ||||||||||||||||||||||||||||||||||||||||
| Iranians | Languages of Iran | Iran | 7/324 | 2.2% | ||||||||||||||||||||||||||||||||||||||||
| Azerbaijani Muslims | Azerbaijani (Turkic) | Uromia | 2/91 | 2.2% | ||||||||||||||||||||||||||||||||||||||||
| Yemenite Jews | Hebrew and Arabic | Yemen | 2/94 | 2.1% | ||||||||||||||||||||||||||||||||||||||||
| Andis | Andi (Northeast Caucasian) | western Dagestan | 1/49 | 2% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Paphos | 2/105 | 1.9% | ||||||||||||||||||||||||||||||||||||||||
| Cypriots | Cypriot Greek | Nicosia | 3/161 | 1.9% | ||||||||||||||||||||||||||||||||||||||||
| Assyrians | Assyrian Neo-Aramaic (Semitic) | Uromia and Tehran | 1/55 | 1.8% | ||||||||||||||||||||||||||||||||||||||||
| Abkhazians | Abkhaz (Northwest Caucasian) | Abkhazia | 1/58 | 1.7% | ||||||||||||||||||||||||||||||||||||||||
| Kuwaitis | Gulf Arabic (Semitic) | Kuwait | 2/117 | 1.7% | vauthors=Triki-Fendri S, Sánchez-Diz P, Rey-González D, Alfadhli S, Ayadi I, Ben Marzoug R, Carracedo Á, Rebai A | title=Genetic structure of the Kuwaiti population revealed by paternal lineages | journal=American Journal of Human Biology | volume=28 | issue=2 | pages=203–12 | year=2016 | pmid=26293354 | doi=10.1002/ajhb.22773 | s2cid=25725954 }} | ||||||||||||||||||||||||||||||
| Greek Orthodox | Koine Greek | Lebanon | 2/116 | 1.7% | ||||||||||||||||||||||||||||||||||||||||
| Mashhad, Razavi Khorasan, Iran | 2/129 | 1.6% | 0.8% T1a3-Y11151 (xY8614) | |||||||||||||||||||||||||||||||||||||||||
| Aeolians | Greek | Smyrna | 1/68 | 1.5% | ||||||||||||||||||||||||||||||||||||||||
| Georgians | Georgian (Kartvelian) | Georgia | 1/66 | 1.5% | ||||||||||||||||||||||||||||||||||||||||
| Turkmens | Turkmen (Oghuz) | Golestan | 1/68 | 1.5% | ||||||||||||||||||||||||||||||||||||||||
| Kumyks | Kumyk (Turkic) | Northern Dagestan | 1/73 | 1.4% | ||||||||||||||||||||||||||||||||||||||||
| Kuban Nogays | Nogai (Turkic) | north of Sea of Azov around Prymorsk | 1/87 | 1.2% | ||||||||||||||||||||||||||||||||||||||||
| Ossetian Digors | Digorian (Scythian) | North Ossetia | 1/127 | 0.8% | ||||||||||||||||||||||||||||||||||||||||
| Yemeni Arabs | Sanaani Arabic (Semitic) | Sana'a | 1/129 | 0.8% | Uta D. Immel et al., "Institut für Rechtsmedizin, Martin-Luther Universität Haale/Saale," "http://www.yhrd.org/" (1999), | |||||||||||||||||||||||||||||||||||||||
| Syrians | Syrian Arabic (Semitic) | Syria | 4/518 | 0.8% | ||||||||||||||||||||||||||||||||||||||||
| Kabardins | Kabardian (Northwest Caucasian) | Kabardino-Balkaria | 1/140 | 0.7% | ||||||||||||||||||||||||||||||||||||||||
| Circassians | Adyghe (Northwest Caucasian) | Republic of Adygea | 1/142 | 0.7% | ||||||||||||||||||||||||||||||||||||||||
| Abkhazians | Abkhaz (Northwest Caucasian) | Abkhazia | 1/162 | 0.6% |
There are also unconfirmed reports of T-M70+ amongst 28% (7/25) of Lezginians in Dagestan, 21.7% (5/23) of Ossetians in Zamankul, 14% (7/50) of Iranians in Isfahan, 13% (3/23) of Ossetians in Zil'ga, 12.6% (11/87) of Kurmanji Kurds in Eastern Turkey, 11.8% (2/17) of Palestinian Arabs in Palestine, 8.3% (1/12) of Iranians in Shiraz, 8.3% (2/24) of Ossetians in Alagir, 8% (2/25) of Kurmanji Kurds in Georgia, 7.5% (6/80) of Iranians in Tehran, 7.4% (10/135) of Palestinian Arabs in Israeli Village, 7% (10/143) of Palestinian Arabs in Israel and Palestine, 5% (1/19) of Chechens in Chechenia, 4.2% (3/72) of Azerbaijanians in Azerbaijan, 4.1% (2/48) of Iranians in Isfahan, 4% (4/100) of Armenians in Armenia, 4% (1/24) of Bedouins in Israel and 2.6% (1/39) of Turks in Ankara.
Africa
Fossils excavated at the Late Neolithic site of Kelif el Boroud in Morocco, which have been radiocarbon-dated to around 3,000 BCE, have been found to belong to haplogroup T-M184.
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Somalis (Dir clan) | Somali (East Cushitic) | Djibouti | 24/24 | 100% | The main sub-clans of the Dir clan in Djibouti are the Issa and Gadabuursi. | ||||||||||||
| Somalis (Dire Dawa) | Somali (East Cushitic) | Dire Dawa | 14/17 | 82.4% | last1=Plaster | display-authors=et al. | year=2011 | title=Variation in Y chromosome, mitochondrial DNA and labels of identity on Ethiopia | journal=UCL Discovery | url=http://discovery.ucl.ac.uk/1331901/3/1331901_CP_Thesis-SUBMITTED-DRAFT-POST-VIVA.pdf}} | Dir sub-clans of Dire Dawa are Issa, Gurgura and Gadabuursi. | ||||||
| Anteony | Antemoro (Plateau Malagasy) | old Antemoro Kingdom | 22/37 | 59.5% | The Anteony are the descendants of aristocrats, from whom the Antemoro king is chosen. Can be grouped into the Silamo, because they have the right to undertake the ritual slaughter of animals (Sombily) | ||||||||||||
| Somalis (Dir clan) and Afars | Somali and Afar (East Cushitic) | Djibouti | 30/54 | 56.6% | Mixed sample of Somali and Afar individuals. | ||||||||||||
| Somalis (Ethiopia) | Somali (East Cushitic) | Shilavo (woreda) (Somali Region of Ethiopia) | 5/10 | 50% | The geographic location of this Ethiopia sample as seen in Fig.1. | ||||||||||||
| Somalis (Isaaq) | Somali (East Cushitic) | Somaliland | 4/4 | 100% | All belonging to the T1a-Y16897 subclade | ||||||||||||
| Afars | Afar language (East Cushitic) | Djibouti | 5/20 | 25% | |||||||||||||
| Toubou | Toubou | Chad | 31% | All belonging to the T1a-PF5662 subclade | |||||||||||||
| Akie | Akie people (Nilotic) | Tanzania | 3/13 | 23.1% | [Hirbo et al.] | Akie people have remnants of a Cushitic language | |||||||||||
| Somalis | Somali (East Cushitic) | Jijiga (Somali Region of Ethiopia) | 19/83 | 22.9% | Jijiga Somalis. | ||||||||||||
| Arabs from Somalia | Somali (East Cushitic) | immigrants in Yemen | 7/33 | 21.2% | vauthors=Immel UD, Kleiber M | title=Y-chromosomal STR haplotypes in an Arab population from Somalia | journal=Forensic Science International: Genetics Supplement Series | volume=2 | issue=1 | year=2009 | pages=409–10 | doi=10.1016/j.fsigss.2009.08.034}} | |||||
| Lemba | Venda and Shona (Bantu) | South Africa | 6/34 | 17.6% | Exclusively belong to T1a2 (old T1b). Possible recent founder effect. Low frequency of T1a2 has been observed in Bulgarian Jews and Turks but is not found in other Jewish communities. Y-str Haplotypes close to some T1a2 Armenians. | ||||||||||||
| Rangi | Rangi Language (Bantu) | Tanzania | 5/32 | 15.6% | [Hirbo et al.] | ||||||||||||
| Multiple ethnicity | - | Somalia | 15/105 | 14.3% | vauthors=Brión M, Sanchez JJ, Balogh K, Thacker C, Blanco-Verea A, Børsting C, Stradmann-Bellinghausen B, Bogus M, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N | title=Introduction of a single nucleodite polymorphism-based 'Major Y-chromosome haplogroup typing kit' suitable for predicting the geographical origin of male lineages | journal=Electrophoresis | volume=26 | issue=23 | pages=4411–20 | year=2005 | pmid=16273584 | doi=10.1002/elps.200500293 | s2cid=25951019 }} | |||
| Iraqw | Iraqw language (Cushitic) | Tanzania | 6/47 | 12.8% | [Hirbo et al.] | ||||||||||||
| Wachagga | Kichagga (Niger-Congo) | Dār as-Salām | 3/24 | 12.5% | vauthors=Xu H, Wang CC, Shrestha R, Wang LX, Zhang M, He Y, Kidd JR, Kidd KK, Jin L, Li H | title=Inferring population structure and demographic history using Y-STR data from worldwide populations | journal=Molecular Genetics and Genomics | volume=290 | issue=1 | pages=141–50 | year=2015 | pmid=25159112 | doi=10.1007/s00438-014-0903-8 | s2cid=15972847 }} | Mixed with Rift Southern Cushites. | ||
| Somali | Somali (Cushitic) | immigrants to Norway | 12/104 | 11.5% | vauthors=Stenersen M, Perchla D, Søvik E, Flønes AG, Dupuy BM | title=Kurdish (Iraq) and Somalian population data for 15 autosomal and 9 Y-chromosomal STR loci | journal=International Congress Series | volume=1261 | year=2004 | pages=185–7 | doi=10.1016/S0531-5131(03)01823-5}} | ||||||
| Bench | Bench(northern Omotic) | Bench Maji Zone | 14/126 | 11.4% | |||||||||||||
| Kores | (Cushitic) | SNNP | 2/18 | 11.1% | |||||||||||||
| Oromo | Afaan Oromo language (Cushitic) | Oromiyaa | 1/9 | 11.1% | vauthors=Wood ET, Stover DA, Ehret C, Destro-Bisol G, Spedini G, McLeod H, Louie L, Bamshad M, Strassmann BI, Soodyall H, Hammer MF | title=Contrasting patterns of Y chromosome and mtDNA variation in Africa: evidence for sex-biased demographic processes | journal=European Journal of Human Genetics | volume=13 | issue=7 | pages=867–76 | year=2005 | pmid=15856073 | doi=10.1038/sj.ejhg.5201408 | doi-access=free }} | |||
| Fulbe | Fula | northern Cameroon | 3/27 | 11.1% | vauthors=Coia V, Brisighelli F, Donati F, Pascali V, Boschi I, Luiselli D, Battaggia C, Batini C, Taglioli L, Cruciani F, Paoli G, Capelli C, Spedini G, Destro-Bisol G | title=A multi-perspective view of genetic variation in Cameroon | journal=American Journal of Physical Anthropology | volume=140 | issue=3 | pages=454–64 | year=2009 | pmid=19425092 | doi=10.1002/ajpa.21088 | bibcode=2009AJPA..140..454C }} | |||
| Gorowa | Gorowa language (Cushitic) | Tanzania | 2/19 | 10.5% | [Hirbo et al.] | ||||||||||||
| Somali | Somali (Cushitic) | immigrants to Denmark | 21/201 | 10.4% | vauthors=Hallenberg C, Simonsen B, Sanchez J, Morling N | title=Y-chromosome STR haplotypes in Somalis | journal=Forensic Science International | volume=151 | issue=2–3 | pages=317–21 | year=2005 | pmid=15939170 | doi=10.1016/j.forsciint.2005.01.011}} | ||||
| Upper Egyptians | Egyptian Arabic | Luxor Governorate | 3/29 | 10.3% | vauthors=Arredi B, Poloni ES, Paracchini S, Zerjal T, Fathallah DM, Makrelouf M, Pascali VL, Novelletto A, Tyler-Smith C | title=A predominantly neolithic origin for Y-chromosomal DNA variation in North Africa | journal=American Journal of Human Genetics | volume=75 | issue=2 | pages=338–45 | year=2004 | pmid=15202071 | pmc=1216069 | doi=10.1086/423147}} | |||
| Kontas | Konta language (Omotic) | Konta special woreda | 11/107 | 10.3% | |||||||||||||
| Rendille | Rendille language (Cushitic) | Marsabit County | 3/31 | 9.7% | [Hirbo et al.] | ||||||||||||
| Datogs | Rendille language (Cushitic) | Tanzania | 3/31 | 9.7% | |||||||||||||
| Gewadas | Gewada language (east Cushitic) | SNNP | 11/116 | 9.5% | |||||||||||||
| Antalaotra | Antemoro (Plateau Malagasy) | old Antemoro Kingdom | 4/43 | 9.3% | The Antalaotra are in charge of the magical and religious domains; they have the ability to read and write Sorabe. Can be grouped into the Silamo, because they have the right to undertake the ritual slaughter of animals (Sombily) | ||||||||||||
| Upper Egyptians | Egyptian Arabic | Aswan Governorate | 1/11 | 9.1% | |||||||||||||
| N'Djamena Mix | Mix | N'Djamena | 5/55 | 9.1% | Marc Haber 2016 | All belonging to the T1a-PF5662 subclade | |||||||||||
| Upper Egyptians | Egyptian Arabic | Assiut Governorate | 6/70 | 8.6% | |||||||||||||
| Konsos | (Semitic) | Konso special woreda | 2/24 | 8.3% | |||||||||||||
| Somali | Somali (Cushitic) | immigrants to Sweden | 12/147 | 8.2% | |||||||||||||
| Arabs and Berbers | Egyptian Arabic and Siwi | Lower Egypt | 12/147 | 8.2% | |||||||||||||
| Upper Egyptians | Egyptian Arabic | Sohag Governorate | 4/52 | 7.7% | |||||||||||||
| Egyptians | Erythraic (Cushitic) | Egypt | 7/92 | 7.6% | If the K* sample is M184+ then 8.7% | ||||||||||||
| Tigrayans | Tigrinya (South Semitic) | Tigray Region | 2/30 | 6.7% | |||||||||||||
| Dirashas | Dirasha (east Cushitic) | Dirashe special woreda | 5/79 | 6.3% | |||||||||||||
| Canarians | Canarian Spanish | Tenerife | 11/178 | 6.2% | |||||||||||||
| Kordofanians | Kordofanian | Kurdufan | 4/69 | 5.8% | |||||||||||||
| Upper Egyptians | Egyptian Arabic | Qena Governorate | 3/52 | 5.8% | |||||||||||||
| Tuareg | Tuareg (Berber) | Gorom-Gorom | 1/18 | 5.6% | vauthors=Pereira L, Cerný V, Cerezo M, Silva NM, Hájek M, Vasíková A, Kujanová M, Brdicka R, Salas A | title=Linking the sub-Saharan and West Eurasian gene pools: maternal and paternal heritage of the Tuareg nomads from the African Sahel | journal=European Journal of Human Genetics | volume=18 | issue=8 | pages=915–23 | year=2010 | pmid=20234393 | pmc=2987384 | doi=10.1038/ejhg.2010.21}} | |||
| Afars | Afar (East Cushitic) | Afar Region | 6/111 | 5.4% | |||||||||||||
| Ethiopians | Ethiopian languages | Ethiopia | 4/74 | 5.4% | |||||||||||||
| Mashiles | Mashile language (Cushitic) | SNNP | 7/130 | 5.4% | |||||||||||||
| Gurages | Gurage languages (South Semitic) | SNNP | 6/118 | 5.1% | |||||||||||||
| Oromo | Afaan Oromo language (Cushitic) | Oromiyaa | 4/78 | 5.1% | |||||||||||||
| Oromo | Afaan Oromo language (Cushitic) | Adis Abeba | 2/40 | 5% | -- | ||||||||||||
| Turu | Nyaturu (Bantu) | Tanzania | 1/20 | 5% | vauthors=Tishkoff SA, Gonder MK, Henn BM, Mortensen H, Knight A, Gignoux C, Fernandopulle N, Lema G, Nyambo TB, Ramakrishnan U, Reed FA, Mountain JL | title=History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation | journal=Molecular Biology and Evolution | volume=24 | issue=10 | pages=2180–95 | year=2007 | pmid=17656633 | doi=10.1093/molbev/msm155 | doi-access=free }} | |||
| Moroccan Jews | Haketia (Romance) | Israel | 1/20 | 5% | vauthors=Cruciani F, La Fratta R, Trombetta B, Santolamazza P, Sellitto D, Colomb EB, Dugoujon JM, Crivellaro F, Benincasa T, Pascone R, Moral P, Watson E, Melegh B, Barbujani G, Fuselli S, Vona G, Zagradisnik B, Assum G, Brdicka R, Kozlov AI, Efremov GD, Coppa A, Novelletto A, Scozzari R | title=Tracing past human male movements in northern/eastern Africa and western Eurasia: new clues from Y-chromosomal haplogroups E-M78 and J-M12 | journal=Molecular Biology and Evolution | volume=24 | issue=6 | pages=1300–11 | year=2007 | pmid=17351267 | doi=10.1093/molbev/msm049 | doi-access=free }} | |||
| Gedeos | Gedeo (east Cushitic) | SNNP | 6/122 | 4.9% | |||||||||||||
| Wairak | Iraqw (Cushitic) | Tanzania | 2/41 | 4.9% | |||||||||||||
| Western Libyans | Libyan Arabic (Semitic) | Tripoli region | 7/142 | 4.9% | vauthors=Triki-Fendri S, Sánchez-Diz P, Rey-González D, Ayadi I, Alfadhli S, Rebai A, Carracedo Á | title=Population genetics of 17 Y-STR markers in West Libya (Tripoli region) | journal=Forensic Science International. Genetics | volume=7 | issue=3 | pages=e59–61 | year=2013 | pmid=23473875 | doi=10.1016/j.fsigen.2013.02.002 | hdl=20.500.11940/4062 | hdl-access=free }} | ||
| Tunisians | Tunisian Arabic (Semitic) | Sfax | 5/105 | 4.8% | vauthors=Ayadi I, Ammar-Keskes L, Rebai A | title=Haplotypes for 13 Y-chromosomal STR loci in South Tunisian population (Sfax region) | journal=Forensic Science International | volume=164 | issue=2–3 | pages=249–53 | year=2006 | pmid=16293385 | doi=10.1016/j.forsciint.2005.10.006}} | ||||
| Libyans | Libyan Arabic (Semitic) | Tripoli area | 3/63 | 4.8% | vauthors=Immel UD, Erhuma M, Mustafa T, Kleiber M, Klintschar M | title=Y-chromosomal STR haplotypes in an Arab population from Libya | journal=International Congress Series | volume=1288 | year=2006 | pages=156–8 | doi=10.1016/j.ics.2005.09.011}} | ||||||
| Kanuri | Kanuri | Cameroon | 1/21 | 4.8% | [Hirbo et al.] | ||||||||||||
| Iraqw | Iraqw (Cushitic) | Tanzania | 2/43 | 4.7% | |||||||||||||
| Yems | Yemsa (Omotic) | SNNP | 5/107 | 4.7% | |||||||||||||
| Jews | (Semitic) | Ethiopia | 1/22 | 4.5% | |||||||||||||
| Gobeze | Cushitic | SNNP | 5/113 | 4.4% | |||||||||||||
| Upper Egyptians | Egyptian Arabic | Minya Governorate | 1/23 | 4.3% | |||||||||||||
| Konsos | Konso language (East Cushitic) | Konso special woreda | 4/94 | 4.3% | |||||||||||||
| Kembaatas | East Cushitic | Kembata Tembaro Zone | 4/102 | 3.9% | |||||||||||||
| Tigrayans | Tigrinya (South Semitic) | Eritrea | 1/28 | 3.6% | |||||||||||||
| Tigrayans | Tigrinya (South Semitic) | Eritrea | 1/31 | 3% | |||||||||||||
| Amharas | Amharic (Semitic) | Ethiopia | 1/34 | 2.9% | |||||||||||||
| Hutus | Rwanda-Rundi (Niger-Congo) | Rwanda | 1/39 | 2.6% | vauthors=Caglià A, Tofanelli S, Coia V, Boschi I, Pescarmona M, Spedini G, Pascali V, Paoli G, Destro-Bisol G | title=A study of Y-chromosome microsatellite variation in sub-Saharan Africa: a comparison between F(ST) and R(ST) genetic distances | journal=Human Biology | volume=75 | issue=3 | pages=313–30 | year=2003 | pmid=14527196 | jstor=41466150 | doi=10.1353/hub.2003.0041 | s2cid=36209595 }} | ||
| Lower Egyptians | Egyptian Arabic (Semitic) | Mansoura | 1/44 | 2.2% | |||||||||||||
| Berbers | Shilha (Berber) | Siwa Oasis | 2/93 | 2.2% | vauthors=Dugoujon JM, Coudray C, Torroni A, Cruciani F, Scozzari R, Moral P, Louali N, Kossman M | chapter=The Berber and the Berbers: Genetic and linguistic diversities | chapter-url= | pages=123–45 | veditors=D'Errico F, Hombart JM | year=2009 | title=Becoming Eloquent: Advances in the Emergence of Language, Human Cognition, and Modern Cultures | publisher=John Benjamins | isbn=978-90-272-3269-4 }} | ||||
| Meru | Meru (Northeast Bantu) | Tanzania | 2/99 | 2% | last1=Charoenchote | first1=Wanwalai | year=2004 | title=AmpFℓSTR Identifiler STR Allele Frequencies and PowerPlex Y-STR Haplotype Frequencies of the Meru Population of Northern Tanzania | publisher=University of Nevada | type=Thesis | oclc=368708609}} | ||||||
| Itam | Ibibio | Obong Itam (Southeast Nigeria) | 1/50 | 2% | vauthors=Veeramah KR, Connell BA, Ansari Pour N, Powell A, Plaster CA, Zeitlyn D, Mendell NR, Weale ME, Bradman N, Thomas MG | title=Little genetic differentiation as assessed by uniparental markers in the presence of substantial language variation in peoples of the Cross River region of Nigeria | journal=BMC Evolutionary Biology | volume=10 | page=92 | year=2010 | issue=1 | pmid=20356404 | pmc=2867817 | doi=10.1186/1471-2148-10-92 | bibcode=2010BMCEE..10...92V | doi-access=free }} | |
| Cape Verdeans | Cape Verdean Creole (Portuguese Creole) | Windward islands São Nicolau, São Vicente, and Santo Antão | 2/101 | 2% | vauthors=Gonçalves R, Rosa A, Freitas A, Fernandes A, Kivisild T, Villems R, Brehm A | title=Y-chromosome lineages in Cabo Verde Islands witness the diverse geographic origin of its first male settlers | journal=Human Genetics | volume=113 | issue=6 | pages=467–72 | year=2003 | pmid=12942365 | doi=10.1007/s00439-003-1007-4 | s2cid=63381583 | hdl=10400.13/3047 | hdl-access=free }} | |
| Ovimbundo | Umbundu and Portuguese | Angola | 1/53 | 1.9% | vauthors=Melo MM, Carvalho M, Lopes V, Anjos MJ, Serra A, Vieira DN, Sequeiros J, Corte-Real F | title=Y-STR haplotypes in three ethnic linguistic groups of Angola population | journal=Forensic Science International. Genetics | volume=5 | issue=3 | pages=e83–8 | year=2011 | pmid=20801729 | doi=10.1016/j.fsigen.2010.08.002}} | ||||
| Tunisians | Tunisian Arabic (Semitic) | Tunis | 1/54 | 1.9% | |||||||||||||
| Berbers | Shilha (Berber) | Asni | 1/54 | 1.9% | |||||||||||||
| Eastern Libyans | Libyan Arabic (Semitic) | Benghazi | 4/214 | 1.9% | vauthors=Elmrghni S, Coulson-Thomas YM, Kaddura M, Dixon RA, Williams DR | title=Population genetic data for 17 Y STR markers from Benghazi (East Libya) | journal=Forensic Science International. Genetics | volume=6 | issue=2 | pages=224–7 | year=2012 | pmid=21640679 | doi=10.1016/j.fsigen.2011.05.001 | url=http://eprints.lincoln.ac.uk/4565/1/FSIGEN_747corrected.pdf }} | |||
| Algerians | Algerian Arabic (Semitic) | Algeria | 3/164 | 1.8% | |||||||||||||
| Baribas | Baatonum (Niger–Congo) | Benin | 1/57 | 1.8% | vauthors=Fortes-Lima C, Brucato N, Croze M, Bellis G, Schiavinato S, Massougbodji A, Migot-Nabias F, Dugoujon JM | title=Genetic population study of Y-chromosome markers in Benin and Ivory Coast ethnic groups | journal=Forensic Science International. Genetics | volume=19 | pages=232–7 | year=2015 | pmid=26275614 | doi=10.1016/j.fsigen.2015.07.021}} | T1a-M70(xT1a2-L131) | ||||
| Bokoras | Karamojong (Eastern Nilotic) | Karamoja region | 1/59 | 1.7% | vauthors=Gomes V, Sánchez-Diz P, Amorim A, Carracedo A, Gusmão L | title=Digging deeper into East African human Y chromosome lineages | journal=Human Genetics | volume=127 | issue=5 | pages=603–13 | year=2010 | pmid=20213473 | doi=10.1007/s00439-010-0808-5 | s2cid=23503728 }} | |||
| Lower Egyptians | Egyptian Arabic (Semitic) | Cairo | 1/63 | 1.6% | vauthors=Manni F, Leonardi P, Barakat A, Rouba H, Heyer E, Klintschar M, McElreavey K, Quintana-Murci L | title=Y-chromosome analysis in Egypt suggests a genetic regional continuity in Northeastern Africa | journal=Human Biology | volume=74 | issue=5 | pages=645–58 | year=2002 | pmid=12495079 | doi=10.1353/hub.2002.0054 | s2cid=26741827 }} | |||
| Tumbuka | Tumbuka (Niger-Congo) | northern Malawi | 1/61 | 1.6% | vauthors=Ansari-Pour N, Moñino Y, Duque C, Gallego N, Bedoya G, Thomas MG, Bradman N | title=Palenque de San Basilio in Colombia: genetic data support an oral history of a paternal ancestry in Congo | journal=Proceedings: Biological Sciences | volume=283 | issue=1827 | article-number=20152980 | year=2016 | pmid=27030413 | pmc=4822459 | doi=10.1098/rspb.2015.2980}} | |||
| Mozabites | Mozabite (Berber) | Ghardaia | 1/68 | 1.5% | vauthors=Bosch E, Calafell F, Pérez-Lezaun A, Comas D, Izaabel H, Akhayat O, Sefiani A, Hariti G, Dugoujon JM, Bertranpetit J | title=Y chromosome STR haplotypes in four populations from northwest Africa | journal=International Journal of Legal Medicine | volume=114 | issue=1–2 | pages=36–40 | year=2000 | pmid=11197625 | doi=10.1007/s004140000136 | s2cid=24279528 }} | |||
| Tunisians | Tunisian Arabic (Semitic) | South Tunisia | 3/200 | 1.5% | vauthors=Makki-Rmida F, Kammoun A, Mahfoudh N, Ayadi A, Gibriel AA, Mallek B, Maalej L, Hammami Z, Maatoug S, Makni H, Masmoudi S | title=Genetic diversity and haplotype structure of 21 Y-STRs, including nine noncore loci, in South Tunisian Population: Forensic relevance | journal=Electrophoresis | volume=36 | issue=23 | pages=2908–13 | year=2015 | pmid=26331800 | doi=10.1002/elps.201500204 | s2cid=866382 }} | |||
| Soussians | Tunisian Arabic (Semitic) | Sousse | 3/220 | 1.4% | vauthors=Fadhlaoui-Zid K, Garcia-Bertrand R, Alfonso-Sánchez MA, Zemni R, Benammar-Elgaaied A, Herrera RJ | title=Sousse: extreme genetic heterogeneity in North Africa | journal=Journal of Human Genetics | volume=60 | issue=1 | pages=41–9 | year=2015 | pmid=25471516 | doi=10.1038/jhg.2014.99 | s2cid=25186140 | doi-access=free }} | ||
| Chewa | Chewa (Niger-Congo) | Malawi | 1/92 | 1.1% | |||||||||||||
| Maasai | Maasai (Eastern Nilotic) | Kinyawa (Mashuru) | 1/100 | 1% | YHRD | ||||||||||||
| Bantu | Narrow Bantu (Niger-Congo) | Pretoria | 1/98 | 1% | |||||||||||||
| Nilotes | Ateker (Eastern Nilotic) | Karamoja region | 1/118 | 0.8% | |||||||||||||
| Andalusians | Andalusian Arabic (Semitic) | Testour, El Alia, Gualaat-El-Andalous, Slouguia | 1/132 | 0.8% | vauthors=Cherni L, Pereira L, Goios A, Loueslati BY, Khodjet el Khil H, Gomes I, Gusmão L, Alves C, Slama A, Amorim A, Elgaaied AB | title=Y-chromosomal STR haplotypes in three ethnic groups and one cosmopolitan population from Tunisia | journal=Forensic Science International | volume=152 | issue=1 | pages=95–9 | year=2005 | pmid=15939181 | doi=10.1016/j.forsciint.2005.02.007}} | Refugees from Al-Andalus following the capitulation of the Islamic kingdoms in Valencia and Granada | |||
| Bantus | Bantu | Botswana, Namibia and Zambia | 1/140 | 0.7% | Father and paternal grandfather belonged to the same ethnolinguistic group | ||||||||||||
| Basothos | Sesotho (Niger-Congo) | Lesotho | 1/181 | 0.6% | vauthors=Montinaro F, Davies J, Capelli C | title=Group membership, geography and shared ancestry: Genetic variation in the Basotho of Lesotho | journal=American Journal of Physical Anthropology | volume=160 | issue=1 | pages=156–61 | year=2016 | pmid=26779678 | doi=10.1002/ajpa.22933 | bibcode=2016AJPA..160..156M | s2cid=32506568 | url=https://ora.ox.ac.uk/objects/uuid:eb47492c-72f4-43f2-aeb8-bc2a276a1d0f }} | |
| Moroccans | Moroccan Arabic (Semitic) | Casablanca metropolitan area | 1/166 | 0.6% | vauthors=Laouina A, El Houate B, Yahia H, Azeddoug H, Boulouiz R, Chbel F | title=Allele frequencies and population data for 17 Y-STR loci (The AmpFlSTR Y-filer) in Casablanca resident population | journal=Forensic Science International. Genetics | volume=5 | issue=1 | pages=e1–3 | year=2011 | pmid=21126935 | doi=10.1016/j.fsigen.2010.10.016}} | The industrial capital of Morocco where the urban growth is maintained by immigration from all parts of Morocco | |||
| Khoisans | Khoisan | Botswana, Namibia and Zambia | 1/371 | 0.3% | vauthors=Barbieri C, Hübner A, Macholdt E, Ni S, Lippold S, Schröder R, Mpoloka SW, Purps J, Roewer L, Stoneking M, Pakendorf B | title=Refining the Y chromosome phylogeny with southern African sequences | journal=Human Genetics | volume=135 | issue=5 | pages=541–53 | year=2016 | pmid=27043341 | pmc=4835522 | doi=10.1007/s00439-016-1651-0}} | Father and paternal grandfather belonged to the same ethnolinguistic group |
South Asia
T1a-M70 in India has been considered to be of West Eurasian origin.
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Kurru | Yerukala (Dravidian) | Andhra Pradesh | 10/18 | 55.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Bauris | Bengali (Indo-Aryan) | West Bengal | 10/19 | 52.6% | K* is found at 6/19, if M70- but M184+, then could be 84.2%. | |||||||||||||||||||||||||||||||||||||||||||||
| Lodha | Lodhi (Sora–Juray–Gorum Munda) | West Bengal | 2/4 | 50% | ||||||||||||||||||||||||||||||||||||||||||||||
| Rajus | Telugu (Dravidian) | Andhra Pradesh | 3/19 | 15.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Maheli | Mahali (Kherwari Munda) | West Bengal | 2/13 | 15.3% | ||||||||||||||||||||||||||||||||||||||||||||||
| Chenchus | Chenchu (Dravidian) | Andhra Pradesh | 3/20 | 15% | K* is found at 7/20, if M70- but M184+, then could be 50% | |||||||||||||||||||||||||||||||||||||||||||||
| Kare Vokkal | Kannada (Dravidian) | Uttara Kannada | 4/30 | 13.3% | doi=10.1016/j.ajhg.2011.05.030 | title=Indian Siddis: African Descendants with Indian Admixture | year=2011 | last1=Shah | first1=Anish M. | last2=Tamang | first2=Rakesh | last3=Moorjani | first3=Priya | last4=Rani | first4=Deepa Selvi | last5=Govindaraj | first5=Periyasamy | last6=Kulkarni | first6=Gururaj | last7=Bhattacharya | first7=Tanmoy | last8=Mustak | first8=Mohammed S. | last9=Bhaskar | first9=L.V.K.S. | last10=Reddy | first10=Alla G. | last11=Gadhvi | first11=Dharmendra | last12=Gai | first12=Pramod B. | last13=Chaubey | first13=Gyaneshwer | last14=Patterson | first14=Nick | last15=Reich | first15=David | last16=Tyler-Smith | first16=Chris | last17=Singh | first17=Lalji | last18=Thangaraj | first18=Kumarasamy | journal=The American Journal of Human Genetics | volume=89 | issue=1 | pages=154–161 | pmid=21741027 | pmc=3135801}} | K* is found at 3/30, if M70- but M184+, then could be 23.3% |
| Banjaras | Lambadi (Indo-Aryan) | Andhra Pradesh | 2/18 | 11.1% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gonds | Gondi (Dravidian) | South Uttar Pradesh | 4/38 | 10.6% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gonds | Gondi (Dravidian) | Madhya Pradesh | 10/139 | 7.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Indians | languages of India | South India | 18/305 | 5.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Maheli | Mahali (Kherwari Munda) | Jamshedpur from Jharkhand; Purulia, Midnapore & other location from West Bengal | 2/38 | 5.3% | vauthors=Kumar V, Reddy AN, Babu JP, Rao TN, Langstieh BT, Thangaraj K, Reddy AG, Singh L, Reddy BM | title=Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations | journal=BMC Evolutionary Biology | volume=7 | article-number=47 | year=2007 | issue=1 | pmid=17389048 | pmc=1851701 | doi=10.1186/1471-2148-7-47 | bibcode=2007BMCEE...7...47K | doi-access=free }} | Two samples from different studies grouped together | |||||||||||||||||||||||||||||||||
| Chenchus | Chenchu (Dravidian) | Andhra Pradesh | 3/61 | 4.9% | Samples from Trivedi et al. and Kivisild et al. | |||||||||||||||||||||||||||||||||||||||||||||
| Banjaras | Lambadi (Indo-Aryan) | Andhra Pradesh | 2/53 | 3.8% | Two samples from different studies grouped together | |||||||||||||||||||||||||||||||||||||||||||||
| Indians | languages of India | East India | 14/367 | 3.8% | ||||||||||||||||||||||||||||||||||||||||||||||
| Gujaratis | Gujarati (Indo-Aryan) | Gujarat | 1/29 | 3.4% | ||||||||||||||||||||||||||||||||||||||||||||||
| Lodha | Lodhi (Sora–Juray–Gorum Munda) | Midnapore & other location from West Bengal | 2/71 | 2.8% | vauthors=Sengupta S, Zhivotovsky LA, King R, Mehdi SQ, Edmonds CA, Chow CE, Lin AA, Mitra M, Sil SK, Ramesh A, Usha Rani MV, Thakur CM, Cavalli-Sforza LL, Majumder PP, Underhill PA | title=Polarity and temporality of high-resolution y-chromosome distributions in India identify both indigenous and exogenous expansions and reveal minor genetic influence of Central Asian pastoralists | journal=American Journal of Human Genetics | volume=78 | issue=2 | pages=202–21 | year=2006 | pmid=16400607 | pmc=1380230 | doi=10.1086/499411 | bibcode=2006AmJHG..78..202S }} | Three samples from different studies grouped together | ||||||||||||||||||||||||||||||||||
| Sahariyas | Saharia (Munda) | Madhya Pradesh | 2/73 | 2.7% | vauthors=Sharma G, Tamang R, Chaudhary R, Singh VK, Shah AM, Anugula S, Rani DS, Reddy AG, Eaaswarkhanth M, Chaubey G, Singh L, Thangaraj K | title=Genetic affinities of the central Indian tribal populations | journal=PLOS ONE | volume=7 | issue=2 | article-number=e32546 | year=2012 | pmid=22393414 | pmc=3290590 | doi=10.1371/journal.pone.0032546 | bibcode=2012PLoSO...732546S | doi-access=free }} | ||||||||||||||||||||||||||||||||||
| Tamtas | (Indo-Aryan) | Bageshwar | 1/34 | 2.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Kshatriyas | (Indo-Aryan) | Pithoragarh | 2/79 | 2.5% | ||||||||||||||||||||||||||||||||||||||||||||||
| Aryas | Arya (Indo-Aryan) | Nainital | 1/46 | 2.2% | ||||||||||||||||||||||||||||||||||||||||||||||
| Laotians | Lao (Tai-Kadai) | Laos | 1/53 | 1.9% | ||||||||||||||||||||||||||||||||||||||||||||||
| Maravars | Tamil (Dravidian) | Ramanathapuram | 1/80 | 1.3% | vauthors=Arunkumar G, Soria-Hernanz DF, Kavitha VJ, Arun VS, Syama A, Ashokan KS, Gandhirajan KT, Vijayakumar K, Narayanan M, Jayalakshmi M, Ziegle JS, Royyuru AK, Parida L, Wells RS, Renfrew C, Schurr TG, Smith CT, Platt DE, Pitchappan R | title=Population differentiation of southern Indian male lineages correlates with agricultural expansions predating the caste system | journal=PLOS ONE | volume=7 | issue=11 | article-number=e50269 | year=2012 | pmid=23209694 | pmc=3508930 | doi=10.1371/journal.pone.0050269 | bibcode=2012PLoSO...750269A | doi-access=free }} | Dry Land Farmers | |||||||||||||||||||||||||||||||||
| Garos | Garo (Sino-Tibetan) | Tangail | 1/120 | 0.8% | vauthors=Hasan M, Momtaz P, Hosen I, Das SA, Akhteruzzaman S | title=Population genetics of 17 Y-chromosomal STRs loci in Garo and Santal tribal populations in Bangladesh | journal=International Journal of Legal Medicine | volume=129 | issue=2 | pages=251–2 | year=2015 | pmid=24577712 | doi=10.1007/s00414-014-0981-5 | s2cid=23031408 }} | Likely P77+ |
With K-M9+, unconfirmed but probable T-M70+: 56.6% (30/53) of Kunabhis in Uttar Kannada, 32.5% (13/40) of Kammas in Andhra Pradesh, 26.8% (11/41) of Brahmins in Visakhapatnam, 25% (1/4) of Kattunaiken in South India, 22.4% (11/49) of Telugus in Andhra Pradesh, 20% (1/5) of Ansari in South Asia, (2/20) of Poroja in Andhra Pradesh, 9.8% (5/51) of Kashmiri Pandits in Kashmir, 8.2% (4/49) of Gujars in Kashmir, 7.7% (1/13) of Siddis (migrants from Ethiopia) in Andhra Pradesh, 5.5% (3/55) of Adi in Northeast India, 5.5% (7/128) of Pardhans in Adilabad, 5.3% (2/38) of Brahmins in Bihar, 4.3% (1/23) of Bagata in Andhra Pradesh, 4.2% (1/24) of Valmiki in Andhra Pradesh, (1/32) of Brahmins in Maharashtra, 3.1% (2/64) of Brahmins in Gujarat, 2.9% (1/35) of Rajput in Uttar Pradesh, 2.3% (1/44) of Brahmins in Peruru, and 1.7% (1/59) of Manghi in Maharashtra.
Also in Desasth-Brahmins in Maharashtra (1/19 or 5.3%) and Chitpavan-Brahmins in Konkan (1/21 or 4.8%), Chitpavan-Brahmins in Konkan (2/66 or 3%).
Central Asia & East Asia
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | |||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Momyns | Old Basmyl/Kazakh (Turkic) | Argyn tribe, Kazakhstan | 6/100 | 6.3% | url=https://web.archive.org/web/20210117175011/http://iggc.kz/wp-content/uploads/2017/03/Rezultaty-raboty-Lab-Pop-Gen-noyab-2016.pdf | date=17 January 2021 }} | The outlier Babasan subclan is excluded from "sample size" and "percentage". 5 out of 6 Clans and 13 out of 19 Subclans have T-M184 members. | ||||||||||||||||||||||||||||||||||||||||||||||
| Meyrams | Old Basmyl/Kazakh (Turkic) | Argyn tribe | 1/10 | 6% | 5 out of 5 Clans and 11 out of 16 Subclans have T-M184 members. | ||||||||||||||||||||||||||||||||||||||||||||||||
| Xibes | Xibe (Tungusic) | Xinjiang, China | 1/8 | 12.5% | vauthors=Cann HM, de Toma C, Cazes L, Legrand MF, Morel V, Piouffre L, Bodmer J, Bodmer WF, Bonne-Tamir B, Cambon-Thomsen A, Chen Z, Chu J, Carcassi C, Contu L, Du R, Excoffier L, Ferrara GB, Friedlaender JS, Groot H, Gurwitz D, Jenkins T, Herrera RJ, Huang X, Kidd J, Kidd KK, Langaney A, Lin AA, Mehdi SQ, Parham P, Piazza A, Pistillo MP, Qian Y, Shu Q, Xu J, Zhu S, Weber JL, Greely HT, Feldman MW, Thomas G, Dausset J, Cavalli-Sforza LL | title=A human genome diversity cell line panel | journal=Science | volume=296 | issue=5566 | pages=261–2 | year=2002 | pmid=11954565 | doi=10.1126/science.296.5566.261b | bibcode=2002Sci...296..261C | s2cid=41595131 }} | ||||||||||||||||||||||||||||||||||||||
| Xibes | Xibe (Tungusic) | Xinjiang | 3/32 | 9.4% | vauthors=Shou WH, Qiao EF, Wei CY, Dong YL, Tan SJ, Shi H, Tang WR, Xiao CJ | title=Y-chromosome distributions among populations in Northwest China identify significant contribution from Central Asian pastoralists and lesser influence of western Eurasians | journal=Journal of Human Genetics | volume=55 | issue=5 | pages=314–22 | year=2010 | pmid=20414255 | doi=10.1038/jhg.2010.30 | doi-access=free }} | |||||||||||||||||||||||||||||||||||||||
| Hans | - | Ili | 3/32 | 9.4% | vauthors=Xue Y, Zerjal T, Bao W, Zhu S, Shu Q, Xu J, Du R, Fu S, Li P, Hurles ME, Yang H, Tyler-Smith C | title=Male demography in East Asia: a north-south contrast in human population expansion times | journal=Genetics | volume=172 | issue=4 | pages=2431–9 | year=2006 | pmid=16489223 | pmc=1456369 | doi=10.1534/genetics.105.054270 | bibcode=2006Genet.172.2431X }} | K* (xNOP) | |||||||||||||||||||||||||||||||||||||
| Bajo sea Nomads | Bajaw (Malayo-Polynesian) | Sulawesi, Indonesia | 2/27 | 7.4% | T1a-M70 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Yugurs | Eastern Yugur and Western Yugur | Sunan Yugur Autonomous County, Gansu, China | 2/32 | 6.3% | K* (xN-M231, O-M175, P-M45) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Tajiks | Tajik (Southwestern Iranian) | Samangan Province, Afghanistan | 1/16 | 6.3% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Khampas | Khams Tibetan (Sino-Tibetan) | Markham | 1/18 | 5.6% | vauthors=Qi X, Cui C, Peng Y, Zhang X, Yang Z, Zhong H, Zhang H, Xiang K, Cao X, Wang Y, Ouzhuluobu, Ouzhuluobu, Ouzhuluobu, Ouzhuluobu, Ouzhuluobu, Wu T, Chen H, Shi H, Su B | title=Genetic evidence of paleolithic colonization and neolithic expansion of modern humans on the tibetan plateau | journal=Molecular Biology and Evolution | volume=30 | issue=8 | pages=1761–78 | year=2013 | pmid=23682168 | doi=10.1093/molbev/mst093 | doi-access=free }} | T-M272 | ||||||||||||||||||||||||||||||||||||||
| Adis | Adi (Sino-Tibetan) | Arunachal Pradesh, India | 3/55 | 5.5% | vauthors=Cordaux R, Weiss G, Saha N, Stoneking M | title=The northeast Indian passageway: a barrier or corridor for human migrations? | journal=Molecular Biology and Evolution | volume=21 | issue=8 | pages=1525–33 | year=2004 | pmid=15128876 | doi=10.1093/molbev/msh151 | doi-access=free }} | |||||||||||||||||||||||||||||||||||||||
| Xibes | Xibe (Tungusic) | (not stated) | 2/41 | 4.9% | K* (xNOP) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Mongolians | Mongolian (Mongolic) | Inner Mongolia, China | 2/45 | 4.4% | K* (xNOP) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Tajiks | Tajik (Southwestern Iranian) | Afghanistan | 2/56 | 3.6% | doi=10.1371/journal.pone.0034288 | title=Afghanistan's Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events | year=2012 | editor1-last=Kayser | editor1-first=Manfred | last1=Haber | first1=Marc | last2=Platt | first2=Daniel E. | last3=Ashrafian Bonab | first3=Maziar | last4=Youhanna | first4=Sonia C. | last5=Soria-Hernanz | first5=David F. | last6=Martínez-Cruz | first6=Begoña | last7=Douaihy | first7=Bouchra | last8=Ghassibe-Sabbagh | first8=Michella | last9=Rafatpanah | first9=Hoshang | last10=Ghanbari | first10=M | last11=Whale | first11=J | last12=Balanovsky | first12=O | last13=Wells | first13=R. S. | last14=Comas | first14=D | last15=Tyler-Smith | first15=C | last16=Zalloua | first16=P. A. | last17=Genographic | first17=Consortium | journal=PLOS ONE | volume=7 | issue=3 | article-number=e34288 | pmid=22470552 | pmc=3314501 | display-authors=8 | bibcode=2012PLoSO...734288H | doi-access=free }} | |
| Uzbeks | Uzbek (Turkic) | Sar-e Pol Province, Afghanistan | 1/28 | 3.6% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Sherpas | Sherpa (Sino-Tibetan) | Khumjung, Namche, Chaurikharka and Lukla | 5/157 | 3.2% | vauthors=Bhandari S, Zhang X, Cui C, Bianba, Liao S, Peng Y, Zhang H, Xiang K, Shi H, Ouzhuluobu, Ouzhuluobu, Ouzhuluobu, Liu S, Gengdeng, Wu T, Qi X, Su B | title=Genetic evidence of a recent Tibetan ancestry to Sherpas in the Himalayan region | journal=Scientific Reports | volume=5 | article-number=16249 | year=2015 | pmid=26538459 | pmc=4633682 | doi=10.1038/srep16249 | bibcode=2015NatSR...516249B }} | K-M9 (xM-P256, NO-M214, P-M45) Parents and grandparents were reported to be Sherpas. Individuals unrelated for at least three generations. | ||||||||||||||||||||||||||||||||||||||
| Oroqen | Oroqen (Tungusic) | (not stated) | 1/31 | 3.2% | K* (xNOP) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Tajiks | Tajik (Southwestern Iranian) | Takhar Province, Afghanistan | 1/35 | 2.9% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Tajiks | Darî (Southwestern Iranian) | Ferghana | 1/35 | 2.9% | vauthors=Balaresque P, Poulet N, Cussat-Blanc S, Gerard P, Quintana-Murci L, Heyer E, Jobling MA | title=Y-chromosome descent clusters and male differential reproductive success: young lineage expansions dominate Asian pastoral nomadic populations | journal=European Journal of Human Genetics | volume=23 | issue=10 | pages=1413–22 | year=2015 | pmid=25585703 | pmc=4430317 | doi=10.1038/ejhg.2014.285}} | |||||||||||||||||||||||||||||||||||||||
| Tibetans | Dbus (Sino-Tibetan) | Dromo, Tibet | 1/39 | 2.6% | T-M272 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Uyghur | Uyghur (Turkic) | Xinjiang | 1/48 (1/4 samples) | 2.1% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Tu | Monguor (Mongolic) | Qinghai, China | 1/50 | 2% | K* (xN-M231, O-M175, P-M45) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Pashtuns | Pashto (Eastern Iranian) | Kunduz Province, Afghanistan | 1/53 | 1.9% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Mongolians | Mongolian (Mongolic) | Mongolia | 1/65 | 1.5% | K* (xNOP) | ||||||||||||||||||||||||||||||||||||||||||||||||
| Kozha Kazakhs (Steppe Clergy) | Kazakh (Turkic) | Kazakhstan | 1/71 | 1.4% | T1a-M70 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Uyghur | Uyghur (Turkic) | Xinjiang | 3/284 | 1.1% | vauthors=Ou X, Wang Y, Liu C, Yang D, Zhang C, Deng S, Sun H | title=Haplotype analysis of the polymorphic 40 Y-STR markers in Chinese populations | journal=Forensic Science International. Genetics | volume=19 | pages=255–62 | year=2015 | pmid=26344901 | doi=10.1016/j.fsigen.2015.08.007}} | |||||||||||||||||||||||||||||||||||||||||
| Uzbeks | Uzbek (Turkic) | Jawzjan Province, Afghanistan | 1/94 | 1.1% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Mongolians | Mongolian (Mongolic) | Inner Mongolia, China | 1/100 | 1% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Ethnic Pashtuns | Pashto (Eastern Iranian) | mainly Kandahar Province, Afghanistan province of | 1/141 | 0.7% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Yousafzai | Pashto (Eastern Iranian) | Khyber Pakhtunkhwa Province, Afghanistan | 1/146 | 0.7% | |||||||||||||||||||||||||||||||||||||||||||||||||
| Uyghur | Uyghur (Turkic) | Hotan Prefecture, Xinjiang, China | 3/478 | 0.6% | author1=TuErXun | author2=NiYaZiBiLiGai | year=2011 | title=Polymorphisms of Y-STRs in Uygur and Kazak Ethnic in Xinjiang | url=http://www.dissertationtopic.net/doc/3012 | publisher=Xinjiang Medical University | type=Thesis}} | ||||||||||||||||||||||||||||||||||||||||||
| Tibetans | Dbus (Sino-Tibetan) | Qüxü, Tibet | 1/203 | 0.5% | T-M272 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Han Chinese | Mandarin (Sino-Tibetan) | Jilin, China | 1/196 | 0.5% | vauthors=Han Y, Li L, Liu X, Chen W, Yang S, Wei L, Xia M, Ma T, Jin L, Li S | title=Genetic analysis of 17 Y-STR loci in Han and Korean populations from Jilin Province, Northeast China | journal=Forensic Science International. Genetics | volume=22 | pages=8–10 | year=2016 | pmid=26799315 | doi=10.1016/j.fsigen.2016.01.003 | doi-access=free }} | ||||||||||||||||||||||||||||||||||||||||
| Mongolians | Mongolian (Mongolic) | Ordos (city), China | 1/258 | 0.4% | vauthors=Gao T, Yun L, Gao S, Gu Y, He W, Luo H, Hou Y | title=Population genetics of 23 Y-STR loci in the Mongolian minority population in Inner Mongolia of China | journal=International Journal of Legal Medicine | volume=130 | issue=6 | pages=1509–1511 | year=2016 | pmid=27515831 | doi=10.1007/s00414-016-1433-1 | s2cid=19999648 }} | Could be 0.8% (2/258) | ||||||||||||||||||||||||||||||||||||||
| Han Chinese | Mandarin (Sino-Tibetan) | Qujing, Yuxi and Honghe County, China | 1/320 | 0.3% | vauthors=Yanmei Y, Tao G, Yubao Z, Chunjie X, Bifeng C, Shi L, Bingying X, Qiang J, Qinyong Z, Wen Z, Shengjun L, Shengjie N | title=Genetic polymorphism of 11 Y-chromosomal STR loci in Yunnan Han Chinese | journal=Forensic Science International. Genetics | volume=4 | issue=2 | pages=e67–9 | year=2010 | pmid=20129460 | doi=10.1016/j.fsigen.2009.06.002}} | K* (xN-M231, O-M175, P-M45) |
Unconfirmed but probable T-M70+: 2% (4/204) of Hui in Liaoning (China), and 0.9% (1/113) of Bidayuh in Sarawak.
Americas (post-colonisation)
| Population | Language | Location | Members/Sample size | Percentage | Source | Notes | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Panchos | Castilian (Romance) | Panchimalco | 3/11 | 27.3% | vauthors=Lovo-Gómez J, Blanco-Verea A, Lareu MV, Brión M, Carracedo A | title=The genetic male legacy from El Salvador | journal=Forensic Science International | volume=171 | issue=2–3 | pages=198–203 | year=2007 | pmid=16916590 | doi=10.1016/j.forsciint.2006.07.005}} | T-M184 | |||||||
| Quechuas | Quechua | Lima Region | 3/11 | 27.3% | Predicted but possible convergence with Q markers. | ||||||||||||||||
| Movimas | Movima language (Language isolate) | Beni | 1/5 | 20% | vauthors=Tirado M, López-Parra AM, Baeza C, Bert F, Corella A, Pérez-Pérez A, Turbón D, Arroyo-Pardo E | title=Y-chromosome haplotypes defined by 17 STRs included in AmpFlSTR Yfiler PCR Amplification Kit in a multi ethnical population from El Beni Department (North Bolivia) | journal=Legal Medicine (Tokyo, Japan) | volume=11 | issue=2 | pages=101–3 | year=2009 | pmid=18974018 | doi=10.1016/j.legalmed.2008.09.002}} | ||||||||
| Colombians | Colombian Spanish (Romance) | Antioquia | 9/51 | 17.6% | vauthors=Rojas W, Parra MV, Campo O, Caro MA, Lopera JG, Arias W, Duque C, Naranjo A, García J, Vergara C, Lopera J, Hernandez E, Valencia A, Caicedo Y, Cuartas M, Gutiérrez J, López S, Ruiz-Linares A, Bedoya G | title=Genetic make up and structure of Colombian populations by means of uniparental and biparental DNA markers | journal=American Journal of Physical Anthropology | volume=143 | issue=1 | pages=13–20 | year=2010 | pmid=20734436 | doi=10.1002/ajpa.21270 | bibcode=2010AJPA..143...13R }} | |||||||
| Colombians | Colombian Spanish (Romance) | Aranzazu, Caldas | 22/190 | 11.6% | vauthors=Rojas W, Campo O, García J, Soto I, Duque C, Bedoya G, Ruiz-Linares A | year=2012 | title=CoanCestría de apellidos y linajes del. Cromosoma y en el noroeste de Colombia: una herramienta útil para establecer migración entre poblaciones | trans-title=Surnames and Y Chromosome Coancestry in northwest Colombia: A useful tool to establish migration between populations | journal=Revista Colombiana de Antropología | volume=48 | issue=1 | pages=49–79 | doi=10.22380/2539472X.890 | doi-access=free }} | |||||||
| Panamanians | Castilian (Romance languages) | Los Santos Province | 3/30 | 10% | |||||||||||||||||
| Centralwest Argentinians | Argentinian Spanish (Romance) | San Luis | 3/30 | 10% | vauthors=Toscanini U, Vullo C, Berardi G, Llull C, Borosky A, Gómez A, Pardo-Seco J, Salas A | title=A comprehensive Y-STR portrait of Argentinean populations | journal=Forensic Science International. Genetics | volume=20 | pages=1–5 | year=2016 | pmid=26433179 | doi=10.1016/j.fsigen.2015.09.002}} | |||||||||
| Colombians | Colombian Spanish (Romance) | Antioquia | 6/61 | 9.8% | Antioquia except Marinilla and its zone of influence | ||||||||||||||||
| Napu runas | Kichwa | Ecuadorian Amazon | 2/21 | 9.5% | vauthors=González-Andrade F, Sánchez D, Martínez-Jarreta B, Budowle B | title=Y-chromosome STR haplotypes in three different population groups from Ecuador (South America) | journal=Journal of Forensic Sciences | volume=53 | issue=2 | pages=512–4 | year=2008 | pmid=18366595 | doi=10.1111/j.1556-4029.2008.00692.x | s2cid=205767262 }} | Predicted but possible convergence with Q markers. | ||||||
| Colombians | Colombian Spanish (Romance) | Soplaviento | 1/11 | 9.1% | T1a-M70 | ||||||||||||||||
| Yanesha | Yanesha | Yurinaqui (Peruvian Amazon) | 1/12 | 8.3% | vauthors=Barbieri C, Heggarty P, Yang Yao D, Ferri G, De Fanti S, Sarno S, Ciani G, Boattini A, Luiselli D, Pettener D | title=Between Andes and Amazon: the genetic profile of the Arawak-speaking Yanesha | journal=American Journal of Physical Anthropology | volume=155 | issue=4 | pages=600–9 | year=2014 | pmid=25229359 | doi=10.1002/ajpa.22616 | bibcode=2014AJPA..155..600B | hdl=11858/00-001M-0000-0024-9E45-D | s2cid=5621046 | hdl-access=free }} | ||||
| Yanesha | Yanesha | Mayme (Peruvian Amazon) | 1/12 | 8.3% | |||||||||||||||||
| Colombians | Colombian Spanish (Romance) | Huila | 3/42 | 7.1% | author=Luz Angela Alonso Morales | title=Caracterización de la población humana de los departamentos de Tolima y Huila. Perspectivas: demográficas, genéticas y socioculturales | type=Master's thesis | publisher=Universidad Nacional de Colombia | year=2013 | url=http://bdigital.unal.edu.co/11272/ | access-date=18 April 2019 | archive-date=18 April 2019 | archive-url=https://web.archive.org/web/20190418050242/http://bdigital.unal.edu.co/11272/ }} | ||||||||
| Bahamians | Bahamian English (West Germanic) | Long Island | 3/43 | 7% | vauthors=Simms TM, Martinez E, Herrera KJ, Wright MR, Perez OA, Hernandez M, Ramirez EC, McCartney Q, Herrera RJ | title=Paternal lineages signal distinct genetic contributions from British Loyalists and continental Africans among different Bahamian islands | journal=American Journal of Physical Anthropology | volume=146 | issue=4 | pages=594–608 | year=2011 | pmid=21989964 | doi=10.1002/ajpa.21616 | bibcode=2011AJPA..146..594S }} | |||||||
| Panamanians | Castilian (Romance languages) | Panama Province | 3/43 | 7% | |||||||||||||||||
| Northwest Argentinians | Argentinian Spanish (Romance) | Mountainous region of San Salvador de Jujuy | 6/86 | 7% | vauthors=Ramallo V, Mucci JM, García A, Muzzio M, Motti JM, Santos MR, Pérez ME, Alfaro EL, Dipierri JE, Demarchi DA, Bravi CM, Bailliet G | title=Comparison of Y-chromosome haplogroup frequencies in eight Provinces of Argentina | journal=Forensic Science International: Genetics Supplement Series | volume=2 | issue=1 | year=2009 | pages=431–2 | doi=10.1016/j.fsigss.2009.08.047}} | |||||||||
| Kolla | Quechua, Aymara and Argentinian Spanish | Mountainous region of Tucumán | 2/29 | 6.9% | vauthors=Toscanini U, Gusmão L, Berardi G, Gomes V, Amorim A, Salas A, Raimondi E | title=Male lineages in South American native groups: evidence of M19 traveling south | journal=American Journal of Physical Anthropology | volume=146 | issue=2 | pages=188–96 | year=2011 | pmid=21826635 | doi=10.1002/ajpa.21562 | bibcode=2011AJPA..146..188T }} | |||||||
| Centralwest Argentinians | Argentinian Spanish (Romance) | Tucumán | 2/30 | 6.7% | |||||||||||||||||
| Guna | Guna (Chibchan languages) | Guna Yala | 1/16 | 6.3% | vauthors=Grugni V, Battaglia V, Perego UA, Raveane A, Lancioni H, Olivieri A, Ferretti L, Woodward SR, Pascale JM, Cooke R, Myres N, Motta J, Torroni A, Achilli A, Semino O | title=Exploring the Y Chromosomal Ancestry of Modern Panamanians | journal=PLOS ONE | volume=10 | issue=12 | article-number=e0144223 | year=2015 | pmid=26636572 | pmc=4670172 | doi=10.1371/journal.pone.0144223 | bibcode=2015PLoSO..1044223G | doi-access=free }} | According to Hamilton 2014, around 2% of Guna people in Guna Yala are Albinos. This is the highest known frequency in the world | ||||
| Basques | Basque (Isolate language) | Nevada | 1/16 | 6.3% | Laura Valverde Potes et al., "Grupo BIOMICs / BIOMICs Research Group," "http://www.yhrd.org/" (2011), | ||||||||||||||||
| Colombians | Colombian Spanish (Romance) | Marinilla, El Peñol, Antioquia, El Santuario, Cocorná, El Carmen de Viboral, Granada, Antioquia and Guatapé | 15/246 | 6.1% | |||||||||||||||||
| Centralwest Argentinians | Argentinian Spanish (Romance) | Mountainous region of La Rioja (Capital) | 5/87 | 5.7% | |||||||||||||||||
| Kolla | Quechua, Aymara and Argentinian Spanish | Mountainous region of Jujuy | 1/18 | 5.6% | doi=10.1016/j.fsigen.2009.08.008 | pmid=20215030 | title=Y-chromosome lineages in native South American population | year=2010 | last1=Blanco-Verea | first1=A. | last2=Jaime | first2=J.C. | last3=Brión | first3=M. | last4=Carracedo | first4=A. | journal=Forensic Science International: Genetics | volume=4 | issue=3 | pages=187–193}} | |
| Colombians | Colombian Spanish (Romance) | Aburrá Valley and Rionegro (Antioquia) | 3/55 | 5.5% | vauthors=Carvajal-Carmona LG, Soto ID, Pineda N, Ortíz-Barrientos D, Duque C, Ospina-Duque J, McCarthy M, Montoya P, Alvarez VM, Bedoya G, Ruiz-Linares A | title=Strong Amerind/white sex bias and a possible Sephardic contribution among the founders of a population in northwest Colombia | journal=American Journal of Human Genetics | volume=67 | issue=5 | pages=1287–95 | year=2000 | pmid=11032790 | pmc=1288568 | doi=10.1016/S0002-9297(07)62956-5}} | |||||||
| Colombians | Colombian Spanish (Romance) | Tolima | 2/41 | 4.9% | |||||||||||||||||
| Venezuelans | Venezuelan Castilian (Romance languages) | Caracas | 3/62 | 4.8% | |||||||||||||||||
| Yanesha | Yanesha | Ñagazu (Peruvian Amazon) | 1/21 | 4.8% | |||||||||||||||||
| Northeast Argentinians | Argentinian Spanish (Romance) | Corrientes | 1/21 | 4.8% | vauthors=Corach D, Lao O, Bobillo C, van Der Gaag K, Zuniga S, Vermeulen M, van Duijn K, Goedbloed M, Vallone PM, Parson W, de Knijff P, Kayser M | title=Inferring continental ancestry of argentineans from Autosomal, Y-chromosomal and mitochondrial DNA | journal=Annals of Human Genetics | volume=74 | issue=1 | pages=65–76 | year=2010 | pmid=20059473 | doi=10.1111/j.1469-1809.2009.00556.x | s2cid=5908692 | hdl=11336/14301 | hdl-access=free }} | |||||
| Colombians | Colombian Spanish (Romance) | Cundinamarca | 1/22 | 4.5% | |||||||||||||||||
| Mestizos | Guatemalan Castilian | Guatemala | 5/115 | 4.4% | vauthors=Martínez-González LJ, Saiz M, Alvarez-Cubero MJ, Gómez-Martín A, Alvarez JC, Martínez-Labarga C, Lorente JA | title=Distribution of Y chromosomal STRs loci in Mayan and Mestizo populations from Guatemala | journal=Forensic Science International. Genetics | volume=6 | issue=1 | pages=136–42 | year=2012 | pmid=21565570 | doi=10.1016/j.fsigen.2011.04.003}} | T-M184 | |||||||
| Northwest Argentinians | Argentinian Spanish (Romance) | Jujuy | 2/50 | 4% | |||||||||||||||||
| Chileans | Chilean Spanish (Romance languages) | Concepción | 8/198 | 4% | vauthors=Toscanini U, Brisighelli F, Moreno F, Pantoja-Astudillo JA, Morales EA, Bustos P, Pardo-Seco J, Salas A | title=Analysis of Y-chromosome STRs in Chile confirms an extensive introgression of European male lineages in urban populations | journal=Forensic Science International. Genetics | volume=21 | pages=76–80 | year=2016 | pmid=26736138 | doi=10.1016/j.fsigen.2015.12.005}} | |||||||||
| Centralwest Argentinians | Argentinian Spanish (Romance) | Mountainous region of Mendoza (Capital) | 3/75 | 4% | |||||||||||||||||
| Mayas | Guatemalan Castilian | Guatemala | 1/110 | 3.6% | T-M184 | ||||||||||||||||
| Yanesha | Yanesha | 7 de Junio - Villa América (Peruvian Amazon) | 1/29 | 3.5% | |||||||||||||||||
| Brazilians | Brazilian Portuguese (Romance) | Serra, Espírito Santo | 1/29 | 3.5% | last1=Raquel | last2=Figueiredo | first2=F. | display-authors = etal | year=2015 | title=Male-specific contributions to the Brazilian population of Espirito Santo | journal=International Journal of Legal Medicine | volume= 130 | issue= 3 | pages=679–681 | doi=10.1007/s00414-015-1214-2 | pmid=26076592 | s2cid=31546021 }} | ||||
| Ecuadorians | Castilian (Romance languages) | Quito | 4/120 | 3.3% | vauthors=Baeza C, Guzmán R, Tirado M, López-Parra AM, Rodríguez T, Mesa MS, Fernández E, Arroyo-Pardo E | title=Population data for 15 Y-chromosome STRs in a population sample from Quito (Ecuador) | journal=Forensic Science International | volume=173 | issue=2–3 | pages=214–9 | year=2007 | pmid=17320323 | doi=10.1016/j.forsciint.2006.09.011 | url=https://durham-repository.worktribe.com/output/1429787 }} | |||||||
| Central Argentinians | Argentinian Spanish (Romance) | La Pampa | 1/30 | 3.3% | |||||||||||||||||
| Central Argentinians | Argentinian Spanish (Romance) | Córdoba | 1/31 | 3.2% | |||||||||||||||||
| Chileans | Chilean Spanish (Romance languages) | Temuco | 6/194 | 3.1% | |||||||||||||||||
| Panamanians | Castilian (Romance languages) | Herrera Province | 1/36 | 2.8% | |||||||||||||||||
| Venezuelans | Venezuelan Castilian (Romance languages) | Maracaibo | 3/111 | 2.7% | |||||||||||||||||
| Chachapoyas | Chacha | northeastern Peruvian Andes | 3/122 | 2.5% | last1=Guevara | first1=Evelyn K. | display-authors = etal | year=2016 | title=MtDNA and Y-chromosomal diversity in the Chachapoya, a population from the northeast Peruvian Andes-Amazon divide | journal=American Journal of Human Biology | volume=28 | issue= 6 | pages=857–867 | doi=10.1002/ajhb.22878 | pmid=27265853 | s2cid=9663568 }} | |||||
| Nicas | Nicaraguan Castilian | Nicaragua | 4/165 | 2.4% | vauthors=Nuñez C, Baeta M, Sosa C, Casalod Y, Ge J, Budowle B, Martínez-Jarreta B | title=Reconstructing the population history of Nicaragua by means of mtDNA, Y-chromosome STRs, and autosomal STR markers | journal=American Journal of Physical Anthropology | volume=143 | issue=4 | pages=591–600 | year=2010 | pmid=20721944 | doi=10.1002/ajpa.21355 | bibcode=2010AJPA..143..591N | s2cid=24849262 }} | Mestizo individuals | |||||
| Colombians | Colombian Spanish (Romance) | Piendamó, Silvia, Puracé, Jambaló, Páez, Popayán, El Tambo, Sotará, La Vega, Cauca, San Sebastián, Cauca and Bolivar | 1/48 | 2.1% | last1=Xavier | first1=Catarina | display-authors = etal | year=2015 | title=Admixture and Genetic Diversity Distribution Patterns of Non-Recombining Lineages of Native American Ancestry in Colombian Populations | journal=PLOS ONE | volume=10 | issue= 3 | article-number=e0120155 | doi=10.1371/journal.pone.0120155 | pmid=25775361 | pmc=4361580 | bibcode=2015PLoSO..1020155X | doi-access=free }} | Mix sample of Ethnicities | ||
| Europeans | Brazilian Portuguese (Romance languages) | Rio Grande do Sul | 5/255 | 2% | |||||||||||||||||
| Chileans | Chilean Spanish (Romance languages) | Santiago de Chile | 4/196 | 2% | |||||||||||||||||
| Centralwest Argentinians | Argentinian Spanish (Romance) | Buenos Aires | 3/150 | 2% | |||||||||||||||||
| Palenques | Palenquero (Castilian-Bantu) | Palenque de San Basilio (Arriba moiety) | 1/52 | 1.9% | |||||||||||||||||
| Quechuas | Quechua | Bolivia | 1/55 | 1.8% | vauthors=Gayà-Vidal M, Moral P, Saenz-Ruales N, Gerbault P, Tonasso L, Villena M, Vasquez R, Bravi CM, Dugoujon JM | title=mtDNA and Y-chromosome diversity in Aymaras and Quechuas from Bolivia: different stories and special genetic traits of the Andean Altiplano populations | journal=American Journal of Physical Anthropology | volume=145 | issue=2 | pages=215–30 | year=2011 | pmid=21469069 | doi=10.1002/ajpa.21487 | bibcode=2011AJPA..145..215G | hdl=11336/94417 | hdl-access=free }} | |||||
| Bahamians | Bahamian English (West Germanic) | Eleuthera | 1/60 | 1.7% | |||||||||||||||||
| Mexicans | Mexican Castilian (Romance languages) | Querétaro | 2/121 | 1.7% | Mestizo individuals | ||||||||||||||||
| Mexicans | Mexican Castilian (Romance languages) | Guanajuato | 1/63 | 1.6% | last1=Santana | last2=Carla | display-authors = etal | year=2014 | title=Genetic Analysis of 17 Y-STRs in a Mestizo Population from the Central Valley of Mexico | journal=Human Biology }} | Mestizo individuals | ||||||||||
| Colombians | Colombian Spanish (Romance) | Peque (Antioquia) | 1/62 | 1.6% | |||||||||||||||||
| Chileans | Chilean Spanish (Romance languages) | Punta Arenas | 3/194 | 1.6% | |||||||||||||||||
| Colombians | Colombian Spanish (Romance) | Cartagena | 1/61 | 1.6% | last1=Noguera | first1=María | display-authors = etal | year=2013 | title=Colombia's racial crucible: Y chromosome evidence from six admixed communities in the Department of Bolivar | journal=Annals of Human Biology | volume=41 | issue= 5 | pages=453–459 | doi=10.3109/03014460.2013.852244 | pmid=24215508 | s2cid=5210893 }} | T1a-M70 | ||||
| Salvadorans | Castilian (Romance) | El Salvador | 2/150 | 1.3% | last1=Monterrosa | first1=Juan Carlos | display-authors = etal | year=2010 | title=Population data for 12 Y-chromosome STR loci in a sample from El Salvador | journal=Legal Medicine | volume=12 | issue= 1 | pages=46–51 | doi=10.1016/j.legalmed.2009.10.003 | pmid=19962926}} | ||||||
| Jamaicans | Jamaican Patois (English creole) | Jamaica | 2/159 | 1.3% | vauthors=Simms TM, Wright MR, Hernandez M, Perez OA, Ramirez EC, Martinez E, Herrera RJ | title=Y-chromosomal diversity in Haiti and Jamaica: contrasting levels of sex-biased gene flow | journal=American Journal of Physical Anthropology | volume=148 | issue=4 | pages=618–31 | year=2012 | pmid=22576450 | doi=10.1002/ajpa.22090}} | ||||||||
| Colombians | Colombian Spanish (Romance) | Cartagena | 2/173 | 1.2% | vauthors=Builes JJ, Martínez B, Gómez A, Caraballo L, Espinal C, Aguirre D, Montoya A, Moreno M, Amorim A, Gusmão L, Bravo ML | title=Y chromosome STR haplotypes in the Caribbean city of Cartagena (Colombia) | journal=Forensic Science International | volume=167 | issue=1 | pages=62–9 | year=2007 | pmid=16455219 | doi=10.1016/j.forsciint.2005.12.015}} | ||||||||
| Panamanians | Castilian (Romance languages) | Chiriquí Province | 1/92 | 1.1% | |||||||||||||||||
| Ticos | Costa Rican Castilian | Costa Rica | 1/100 | 1% | last1=Villalta | first1=M. | display-authors = etal | year=2008 | title=Haplotype data for 12 Y-chromosome STR loci from Costa Rica | journal=Forensic Science International: Genetics Supplement Series | volume=1 | pages=252–254 | doi=10.1016/j.fsigss.2007.10.101 | doi-access=free }} | |||||||
| Brazilians | Brazilian Portuguese (Romance) | Santa Catarina | 1/109 | 0.9% | last1=Cainé | first1=Laura M. | display-authors = etal | year=2010 | title=Y-chromosomal STR haplotype diversity in males from Santa Catarina, Brazil | journal=Journal of Forensic and Legal Medicine | volume=17 | issue= 2 | pages=92–95 | doi=10.1016/j.jflm.2009.07.023 | pmid=20129429 }} | ||||||
| Virgin islanders | Virgin Islands Creole English (Germanic) | Saint Thomas (Virgin Islands) | 1/134 | 0.8% | vauthors=Benn Torres J, Kittles RA, Stone AC | title=Mitochondrial and Y chromosome diversity in the English-speaking Caribbean | journal=Annals of Human Genetics | volume=71 | issue=Pt 6 | pages=782–90 | year=2007 | pmid=17596204 | doi=10.1111/j.1469-1809.2007.00380.x | s2cid=8723477 }} | |||||||
| Hondurans | Honduran Castilian | Honduras | 1/128 | 0.8% | last1=Matamoros | first1=Mireya | display-authors = etal | year=2009 | title=Population data for 12 Y-chromosome STR loci in a sample from Honduras | journal=Legal Medicine | volume=11 | issue= 5 | pages=251–255 | doi=10.1016/j.legalmed.2009.06.001 | pmid=19628418}} | Mestizo individuals | |||||
| Admixed population | - | Macapá | 1/138 | 0.7% | last1=Abdon da Costa Francez | first1=Pablo | display-authors = etal | year=2012 | title=Haplotype diversity of 17 Y-str loci in an admixed population from the Brazilian Amazon | journal=Genetics and Molecular Biology | volume= 35 | issue= 1 | pages= 45–52 | doi=10.1590/s1415-47572011005000061 | pmid=22481873 | pmc=3313515}} | |||||
| Belizeans | Belizean Castilian and Belizean Creole | Belize | 1/157 | 0.6% | last1=Flores | first1=Shahida | display-authors = etal | year=2015 | title=Allele frequencies for 15 autosomal STR loci and haplotype data for 17 Y-STR loci in a population from Belize | journal=International Journal of Legal Medicine | volume=129 | issue= 6 | pages=1217–1218 | doi=10.1007/s00414-014-1082-1 | pmid=25193820 | s2cid=149715 }} | |||||
| Chileans | Chilean Spanish (Romance languages) | Iquique | 1/207 | 0.5% | |||||||||||||||||
| Brazilians | Brazilian Portuguese (Romance) | Espírito Santo | 1/253 | 0.4% | last1=Figueiredo | first1=Raquel de F. | display-authors = etal | year=2016 | title=Male-specific contributions to the Brazilian population of Espirito Santo | journal=International Journal of Legal Medicine | volume= 130 | issue= 3 | pages= 679–681 | doi=10.1007/s00414-015-1214-2 | pmid=26076592 | s2cid=31546021 }} |
Ancient DNA
Abel Beth Maacah
Abel Beth Maacah 2201 was a man with Y-DNA T-CTS2860 who lived between 1014 - 836 BCE during the Levant Iron Age and was found in the region now known as Abel Beth Maacah, Metula,Israel . At the Iron Age layer which also produces a Yahwistic inscription on a pottery jar from the biblical site of Abel-beth-maachah, which bears a faint Hebrew inscription of the name "Benayau". I2201 from Agranat-Tamir et al 2020.
Ancient Egypt
Egyptian mummy 2516 was a man who lived between 798 - 591 BCE during the Third Intermediate Age and was found in the region now known as Egypt. He is wearing a curly wig, a shabti made of multicoloured wood and a multicoloured wesekh-collar. There is an inscription, encircling the entire body in horizontal lines, with the text of Chapter VI of the Book of the Dead. The Ancient Egyptian was under T-Y6671, the saharan offshoot of T-L208 ultimately derived from T-M70. The finding was by Wurst et al 2024.
Ancient Nubia
Multiple Nubians from Kulubnarti site were found to be of the Haplogroup T lineage (T-Y6671), same as the ancient Egyptian clade. The Kulubnarti Nubians had ~43% Nilotic-related ancestry (individual variation between ~36–54%) with the remaining ancestry consistent with being introduced through Egypt and ultimately deriving from an ancestry pool like that found in the Bronze and Iron Age Levant. It is hypothesized "T" lineage originated or evolved in the Levant, and became Saharan Pastoralists via their spread into Africa during the Neolithic. T-Y6671 is associated with this spread. This falls in line perfectly when considering the Levantine-like DNA that the Nubians harbor in concomitance to T-Y6671. The Nubian samples include I6328, I6340 & I19140. These Nubians lived during 700 - 990 CE and were found in "R and S Cemeteries", where E & J haplogroup was buried amongst these "T" individuals. The finding is presented by Sirak et al.
Neolithic North Africans
During the Neolithic Era a new ancestry from the Levant appears in the Maghreb, coinciding with the arrival of pastoralism in the region, and all three ancestries blend together during the Late Neolithic. This places Haplogroup T as a pastoralist lineage, and due to its circumstances, is associated with Levantine expansion,spreading Afro-Asiatic languages, eventually morphing into Saharan Pastoralists and spreading Afro-Asiatic languages. Sample SKH003 and SKH002 were Neolithic local Northwest African (Maghrebis) and differentiated from older Northwest Africans exactly due to an influx of Levantine PPNB ancestry. This ancestry was introduced with this new Y-chromosome haplogroup (T-M70), and is very clearly a Male dominated migration, as only Y-chromosome lineages were replaced, and no mtdna was introduced. Unlike the earlier expansion of Anatolian Neolithic / Early European farmer dna, which were maternally lead migrations. "Because this Neolithic Levantine ancestry has not been observed on the European side of the Mediterranean during the Neolithic, it probably represents an independent expansion of people from the Levant into North Africa."
Peki'in Cave, Israel
A 2018 study conducted by scholars from Tel-Aviv University, the Israel Antiquities Authority and Harvard University had discovered that 22 out of the 600 people who were buried in Peki'in cave from the Chalcolithic Period were of both local Levantine and Zagros area ancestries, or as phrased in the paper itself: "Ancient DNA from Chalcolithic Israel reveals the role of population mixture in cultural transformation," the scientists concluded that the homogeneous community found in the cave could source ~57% of its ancestry from groups related to those of the local Levant Neolithic, ~26% from groups related to those of the Anatolian Neolithic, and ~17% from groups related to those of the Iran Chalcolithic.". The scholars noted that the Zagros genetic material held "Certain characteristics, such as genetic mutations contributing to blue eye color, were not seen in the DNA test results of earlier Levantine human remains MTDNA blue-eyed, fair-skinned community didn't continue, but at least now researchers have an idea why. "These findings suggest that the rise and fall of the Chalcolithic culture are probably due to demographic changes in the region".
We find that the individuals buried in Peqi'in Cave represent a relatively genetically homogenous population. This homogeneity is evident not only in the genome-wide analyses but also in the fact that most of the male individuals (nine out of ten) belong to the Y-chromosome Haplogroup T (Y-DNA), a lineage thought to have diversified in the Near East. This finding contrasts with both earlier (Neolithic and Epipaleolithic) Levantine populations, which were dominated by Haplogroup E (Y-DNA), and later Bronze Age individuals, all of whom belonged to Haplogroup J (Y-DNA).
Ancient city of Ebla
In the ancient city of Ebla in Syria in the Bronze Age, one individual was found belonging to haplogroup T-L162 (T1a1).
Alalakh Amorite city-state
One individual from Alalakh who lived circa 2014-1781 BC, belonged to haplogroup T-CTS11451 (T1a1a).
Notable haplogroup members
Elite endurance runners
Possible patterns between Y-chromosome and elite endurance runners were studied in an attempt to find a genetic explanation to the Ethiopian endurance running success. Given the superiority of East African athletes in international distance running over the past four decades, it has been speculated that they are genetically advantaged. Elite marathon runners from Ethiopia were analysed for K*(xP) which according to the previously published Ethiopian studies is attributable to the haplogroup T.
According to further studies, T1a1a* (L208) was found to be proportionately more frequent in the elite marathon runners sample than in the control samples than any other haplogroup, therefore this y-chromosome could play a significant role in determining Ethiopian endurance running success. Haplogroup T1a1a* was found in 14% of the elite marathon runners sample of whom 43% of this sample are from Arsi province. In addition, haplogroup T1a1a* was found in only 4% of the Ethiopian control sample and only 1% of the Arsi province control sample. T1a1a* is positively associated with aspects of endurance running, whereas E1b1b1 (old E3b1) is negatively associated.
House of Khalifa
The ruling family of the Kingdom of Bahrain is the House of Khalifa (Arabic: آل خليفة, romanized: Āl Khalīfah) is confirmed West Asian Y-DNA Haplogroup T-L206 subclade of P77*.
The house belongs to the Utab tribe, which is part of the larger Anizah tribal confederation, that migrated from Central Arabia to Kuwait and then ruled all of Qatar. In 1999, Hamad bin Isa Al Khalifa became the Emir of Bahrain and proclaimed himself the King of Bahrain in 2002.
The T-FT364053 haplogroup of the house was determined by DNA testing of descendants in the T-Arab Y DNA Haplogroup Project on Family Tree DNA and other Arab world projects.
Thomas Jefferson
A notable member of the T-M184 haplogroup is American President Thomas Jefferson (most distant known ancestor "MDKA" is Samuel Jefferson, Born 11 October 1607 in Pettistree, Suffolk, England). The Y-chromosomal complement of the Jefferson male line was studied in 1998 in an attempt to resolve the controversy over whether he had fathered the mixed-race children of his slave Sally Hemings. A 1998 DNA study of the Y chromosome in the Jefferson male line found that it matched that of a descendant of Eston Hemings, Sally Hemings' youngest son. This confirmed the body of historical evidence, and most historians believe that Jefferson had a long-term intimate liaison with Hemings for 38 years, and fathered her six children of record, four of whom lived to adulthood. In addition, the testing conclusively disproved any connection between the Hemings descendant and the Carr male line. Jefferson grandchildren had asserted in the 19th century that a Carr nephew had been the father of Hemings' children, and this had been the basis of historians' denial for 180 years. Jefferson's paternal family traced back Wales, where T is incredibly rare, as it is less than
Family Tree DNA, found that the Jefferson T patrilineage belongs to T-BY78550 a subclade of T-PF7444 which is likely of MENA Middle Eastern North African Origins. Spencer Wells who led The Genographic Project places his origin to Canaan
Phylogenetic tree
| {{cladogram | title= Phylogenetic tree of haplogroup T-M184 & closely related macro-lineages | caption= | style= background: | label1=LT | ||||
|---|---|---|---|---|---|---|---|---|
| L298 (43,900 BP) | 1={{clade | LT*]] (basal subclade) | 1=(LTxM184, M20; all cases without M184 or M20.) | label2=TM184 (39,300-45,100 BP) | thickness=3 | label1=T*(xL206) | 1=All cases without L206 or PH110 | label2= |
| T1L206 (26,600 BP) | thickness=3 | label1=T1*(xM70) | 1= | label2= | ||||
| T1aM70 (19,000-30,000 BP) | thickness=3 | label1=T1a*(xL162, L131, Y11151) | 1=All cases without L162, L131 or Y11151 | label2= | ||||
| T1a1L162 (15,400 BP) | thickness=3 | label1=T1a1*(xL208) | 1= | label2= | ||||
| T1a1aL208 (14,800 BP) | thickness=3 | label1=T1a1a*(xCTS11451, Y16897) | 1=All cases without CTS11451 or Y16897 | label2= | ||||
| T1a1a1CTS11451 (9,500 BP) | thickness=3 | label1=T1a1a1*(xY4119, Y6671) | 1=All cases without Y4119 or Y6671 | label2= | ||||
| T1a1a1aY4119 (9,200 BP) | thickness=3 | label1=T1a1a1a*(xCTS2214) | 1=All cases without CTS2214 | label2= | ||||
| T1a1a1a1CTS2214 (8,900 BP) | thickness=3}} | 3= | label3= | |||||
| T1a1a1a2Y6671 (8,900 BP)}} | 3= | label3= | ||||||
| T1a1a1bY6671 (9,200 BP)}} | 3= | label3= | ||||||
| T1a1a2Y16897 (9,500 BP)}}}} | 3= | label3= | ||||||
| T1a2L131 (15,400 BP) | 4= | label4= | ||||||
| T1a3Y11151 (15,400 BP) | label3=T2 | |||||||
| PH110 (26,600 BP) | 3= | L]]''' | ||||||
| M20 | thickness=1 | label1=L1 | ||||||
| M22 | 1=(Mostly South Asia and Central Asia.) | label2= | ||||||
| L2 | ||||||||
| L595 | 2=(The highest diversity and incidence of this rare lineage is found in Europe.) |
Nomenclatural history
Main article: Conversion table for Y chromosome haplogroups
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 | ISOGG 2013 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T-M184 | 26 | VIII | 1U | 25 | Eu16 | H5 | F | K* | K | T | T | K2 | K2 | T | T | T | T | T | T |
| K-M70/T-M70 | 26 | VIII | 1U | 25 | Eu15 | H5 | F | K2 | K2 | T | T1 | K2 | K2 | T | T | T | T1 | T1a | T1a |
| T-P77 | 26 | VIII | 1U | 25 | Eu15 | H5 | F | K2 | K2 | T2 | T1a2 | K2 | K2 | T2 | T2 | T2a1 | T1a1b | T1a1a1 | T1a1a1 |
Original research publications
The following research teams per their publications were represented in the creation of the YCC Tree.
α and
β
γ
δ
ε
ζ
η
Y-DNA backbone tree
Notes
References
Original research
Other works cited
Sources for conversion tables
References
- "T YTree".
- Other SNPs – ''M272'', ''PAGES129'', ''L810'', ''L455'', ''L452'', and ''L445'' – are considered to be [[phylogenetic]]ally equivalent to ''M184''.
- "ISOGG 2018 Y-DNA Haplogroup T".
- (2011). "Increased resolution of Y chromosome haplogroup T defines relationships among populations of the Near East, Europe, and Africa". Human Biology.
- Michael Hodd, ''East Africa Handbook'', 7th Edition, (Passport Books: 2002), p. 21: "To the north are the countries of the Horn of Africa comprising Somaliland
- "The Republic of Somaliland's Position on the Somaliland - Somalia Talks".
- (4 April 2023). "Somaliland and the Need to Update International Law on Statehood Recognition".
- Giuseppe Iacovacci ''et al.'', "Forensic data and microvariant sequence characterization of 27 Y-STR loci analyzed in four Eastern African countries," ''^Forensic Science International: Genetics'', 2016
- "T-Y16897 YTree".
- (2008). "Two sources of the Russian patrilineal heritage in their Eurasian context". American Journal of Human Genetics.
- (2006). "Data for Y-chromosome haplotypes defined by 17 STRs (AmpFLSTR Yfiler) in two Tunisian Berber communities". Forensic Science International.
- (2011). "Genetic data for 17 Y-chromosomal STR loci in Macedonians in the Republic of Macedonia". Forensic Science International. Genetics.
- (2016). "Genomic insights into the origin of farming in the ancient Near East". Nature.
- (2006). "Differential maternal and paternal contributions to the genetic pool of Ibiza Island, Balearic Archipelago". American Journal of Physical Anthropology.
- (2008). "Identifying genetic traces of historical expansions: Phoenician footprints in the Mediterranean". American Journal of Human Genetics.
- (2008). "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula". The American Journal of Human Genetics.
- (2009). "Genetic sub-structure in western Mediterranean populations revealed by 12 Y-chromosome STR loci". International Journal of Legal Medicine.
- (2014). "Genetic diversity in Puerto Rico and its implications for the peopling of the Island and the West Indies". American Journal of Physical Anthropology.
- (2008). "The distribution of Y-chromosome STRs in Dominican population". Forensic Science International: Genetics Supplement Series.
- (2009). "Y-STR variation among ethnic groups from Ecuador: Mestizos, Kichwas, Afro-Ecuadorians and Waoranis". Forensic Science International. Genetics.
- (2008). "Usefulness of 12 Y-STRs for forensic genetics evaluation in two populations from Venezuela". Legal Medicine.
- (2009). "Y-chromosome haplotype database in Venezuelan central region and its comparison with other Venezuelan populations". Forensic Science International: Genetics Supplement Series.
- (2006). "Y-chromosome STRs in an Antioquian (Colombia) population sample". Forensic Science International.
- (1999). "Haplotype frequencies of eight Y-chromosome STR loci in Barcelona (North-East Spain)". International Journal of Legal Medicine.
- (2009). "Population data of 17 Y-STR loci from Rio Grande do Sul state (South Brazil)". Forensic Science International. Genetics.
- (2015). "The Y-chromosome tree bursts into leaf: 13,000 high-confidence SNPs covering the majority of known clades". Molecular Biology and Evolution.
- (2007). "[Gene pool differences between northern and southern Altaians inferred from the data on Y-chromosomal haplogroups]". Genetika.
- (2013). "Introducing the Algerian mitochondrial DNA and Y-chromosome profiles into the North African landscape". PLOS ONE.
- (2012). "The Caucasus as an asymmetric semipermeable barrier to ancient human migrations". Molecular Biology and Evolution.
- (2009). "In search of the pre- and post-neolithic genetic substrates in Iberia: evidence from Y-chromosome in Pyrenean populations". Annals of Human Genetics.
- (2009). "Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation". BMC Evolutionary Biology.
- (2012). "Haplotype data for 16 Y-chromosome STR loci in Aboriginal and Caucasian populations in South Australia". Forensic Science International. Genetics.
- (2011). "Y-chromosome variation in Altaian Kazakhs reveals a common paternal gene pool for Kazakhs and the influence of Mongolian expansions". PLOS ONE.
- (2016). "Y chromosome haplotype diversity in Mongolic-speaking populations and gene conversion at the duplicated STR DYS385a,b in haplogroup C3-M407". Journal of Human Genetics.
- (2013). "Genetic population study of 11 Y chromosome STR loci in Greece". Forensic Science International. Genetics.
- (2015). "Development of an Italian RM Y-STR haplotype database: Results of the 2013 GEFI collaborative exercise". Forensic Science International. Genetics.
- (2004). "Y-chromosomal STR haplotypes in a population sample from continental Greece, and the islands of Crete and Chios". Forensic Science International.
- (2006). "Genetic structure in contemporary south Tyrolean isolated populations revealed by analysis of Y-chromosome, mtDNA, and Alu polymorphisms". Human Biology.
- (2003). "Multiple origins of Ashkenazi Levites: Y chromosome evidence for both Near Eastern and European ancestries". American Journal of Human Genetics.
- (2006). "Y-chromosomal STR haplotypes in a Northeast Italian population sample using 17plex loci PCR assay". International Journal of Legal Medicine.
- (2013). "Uniparental markers in Italy reveal a sex-biased genetic structure and different historical strata". PLOS ONE.
- (2009). "Differential Greek and northern African migrations to Sicily are supported by genetic evidence from the Y chromosome". European Journal of Human Genetics.
- (March 2010). "Phylogeographic analysis of paternal lineages in NE Portuguese Jewish communities". Am. J. Phys. Anthropol..
- (2015). "Portuguese crypto-Jews: the genetic heritage of a complex history". Frontiers in Genetics.
- (2015). "Commentary: Portuguese crypto-Jews: the genetic heritage of a complex history". Frontiers in Genetics.
- (2013). "Y-chromosome polymorphisms and ethnic group - a combined STR and SNP approach in a population sample from northern Italy". Croatian Medical Journal.
- (2001). "Human Y-chromosome variation in the western Mediterranean area: implications for the peopling of the region". Human Immunology.
- (2015). "Traces of forgotten historical events in mountain communities in Central Italy: A genetic insight". American Journal of Human Biology.
- (2012). "Evidence of pre-Roman tribal genetic structure in Basques from uniparentally inherited markers". Molecular Biology and Evolution.
- (2004). "Reduced genetic structure of the Iberian peninsula revealed by Y-chromosome analysis: implications for population demography". European Journal of Human Genetics.
- (2012). "Leonese dialects in Portugal: linguistic-genetic relationships through Y chromosome analysis". Universidade do Porto.
- (2016). "Y chromosome diversity in a linguistic isolate (Mirandese, NE Portugal)". American Journal of Human Biology.
- (2016). "The relationship between surname frequency and Y chromosome variation in Spain". European Journal of Human Genetics.
- (2012). "Assessing the genetic influence of ancient sociopolitical structure: micro-differentiation patterns in the population of Asturias (Northern Spain)". PLOS ONE.
- (2014). "Mitochondrial DNA and Y-chromosome structure at the Mediterranean and Atlantic façades of the Iberian Peninsula". American Journal of Human Biology.
- (2007). "Paleolithic Y-haplogroup heritage predominates in a Cretan highland plateau". European Journal of Human Genetics.
- (2007). "Y chromosome genetic variation in the Italian peninsula is clinal and supports an admixture model for the Mesolithic-Neolithic encounter". Molecular Phylogenetics and Evolution.
- (2003). "Y Chromosome and Mitochondrial DNA Characterization of Pasiegos, a Human Isolate from Cantabria (Spain)". Annals of Human Genetics.
- (2012). "Patterns of Y-STR variation in Italy". Forensic Science International. Genetics.
- (2016). "Genetic heritage of Croatians in the Southeastern European gene pool-Y chromosome analysis of the Croatian continental and Island population". American Journal of Human Biology.
- (2006). "Micro-phylogeographic and demographic history of Portuguese male lineages". Annals of Human Genetics.
- (2005). "Y-chromosome short tandem repeat (STR) haplotypes in a Campania population sample". Clinical Chemistry and Laboratory Medicine.
- Decorte R. ''et al.'', ''YHRD''
- (2013). "Demographic histories, isolation and social factors as determinants of the genetic structure of Alpine linguistic groups". PLOS ONE.
- (2012). "Y-STR genetic diversity in autochthonous Andalusians from Huelva and Granada provinces (Spain)". Forensic Science International. Genetics.
- (2002). "Y-chromosomal STR haplotypes in an Albanian population sample". Forensic Science International.
- (2013). "Y-chromosome diversity in modern Bulgarians: new clues about their ancestry". PLOS ONE.
- (2016). "Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences". Nature Genetics.
- (2010). "Drawing the history of the Hutterite population on a genetic landscape: inference from Y-chromosome and mtDNA genotypes". European Journal of Human Genetics.
- (2008). "Y-chromosome based evidence for pre-neolithic origin of the genetically homogeneous but diverse Sardinian population: inference for association scans". PLOS ONE.
- (2015). "Y-chromosome diversity in Catalan surname samples: insights into surname origin and frequency". European Journal of Human Genetics.
- (2015). "Genetic Heritage of the Balto-Slavic Speaking Populations: A Synthesis of Autosomal, Mitochondrial and Y-Chromosomal Data". PLOS ONE.
- (2009). "Population data for 15 autosomal STRs loci and 12 Y chromosome STRs loci in a population sample from the Sardinia island (Italy)". Legal Medicine.
- (2005). "Analysis of Y-chromosome variability and its comparison with mtDNA variability reveals different demographic histories between islands in the Azores Archipelago (Portugal)". Annals of Human Genetics.
- Carolina Nuñez ''et al.'', Highly discriminatory capacity of the PowerPlex Y23 System for the study of isolated populations 2015.
- (2015). "The Greeks in the West: genetic signatures of the Hellenic colonisation in southern Italy and Sicily". European Journal of Human Genetics.
- (2014). "A global analysis of Y-chromosomal haplotype diversity for 23 STR loci". Forensic Science International. Genetics.
- (2005). "Y-chromosomal STR haplotypes in Macedonian population samples". Forensic Science International.
- (2010). "Assembly of a large Y-STR haplotype database for the Czech population and investigation of its substructure". Forensic Science International. Genetics.
- (2012). "Temporal differentiation across a West-European Y-chromosomal cline: genealogy as a tool in human population genetics". European Journal of Human Genetics.
- (2012). "Analysis of a genetic isolate: the case of Carloforte (Italy)". Human Biology.
- (2015). "High Y-chromosomal diversity and low relatedness between paternal lineages on a communal scale in the Western European Low Countries during the surname establishment". Heredity.
- (2013). "Low-pass DNA sequencing of 1200 Sardinians reconstructs European Y-chromosome phylogeny". Science.
- (2015). "Non-random distribution of 17 Y-chromosome STR loci in different areas of Sardinia". Forensic Science International. Genetics.
- (2010). "STR genetic diversity in a Mediterranean population from the south of the Iberian Peninsula". Annals of Human Biology.
- (2010). "Y Chromosome Single Nucleotide Polymorphisms Typing by SNaPshot MINISEQUENCING". Balkan Journal of Medical Genetics.
- (April 2017). "Estudi genètic dels cognoms catalans, valencians i balears". Csic-Upf }}{{unreliable source?.
- (2011). "Paternal Genetic History of the Basque Population of Spain". Human Biology.
- (2005). "Allelverteilung Y-chromosomaler Short Tandem Repeats in Vorpommern". Greifswald Universitätsbibliothek.
- (2013). "Uniparental genetic heritage of belarusians: encounter of rare middle eastern matrilineages with a central European mitochondrial DNA pool". PLOS ONE.
- (2004). "The origin of the isolated population of the Faroe Islands investigated using Y chromosomal markers". Human Genetics.
- (2011). "Y-chromosomal diversity of the Valachs from the Czech Republic: model for isolated population in Central Europe". Croatian Medical Journal.
- (2016). "Yfiler Plus amplification kit validation and calculation of forensic parameters for two Austrian populations". Forensic Science International. Genetics.
- (2013). "Contemporary paternal genetic landscape of Polish and German populations: from early medieval Slavic expansion to post-World War II resettlements". European Journal of Human Genetics.
- (2017). "Micro and macro geographical analysis of Y-chromosome lineages in South Iberia". Forensic Science International. Genetics.
- (2000). "Y-chromosome variation and Irish origins". Nature.
- (2005). "Y chromosome polymorphisms and haplotypes in South Saxony-Anhalt (Germany)". Forensic Science International.
- (2007). "Haplotype frequencies of 16 Y-chromosome STR loci in the Barcelona metropolitan area population using Y-Filer kit". Forensic Science International.
- (2004). "Differentiation of mitochondrial DNA and Y chromosomes in Russian populations". Human Biology.
- F. Di Giacomo. (2003). "Clinal patterns of human Y chromosomal diversity in continental Italy and Greece are dominated by drift and founder effects". Molecular Phylogenetics and Evolution.
- (2007). "Y chromosome haplotypes in Central-South Italy: implication for reference database". Forensic Science International.
- (2005). "High-resolution phylogenetic analysis of southeastern Europe traces major episodes of paternal gene flow among Slavic populations". Molecular Biology and Evolution.
- (2008). "Genetic origin, admixture, and asymmetry in maternal and paternal human lineages in Cuba". BMC Evolutionary Biology.
- (2006). "Population data for Y-chromosome STR haplotypes from Piedmont (Italy)". Forensic Science International.
- (2008). "Boundaries and clines in the West Eurasian Y-chromosome landscape: insights from the European part of Russia". American Journal of Physical Anthropology.
- (2006). "A 9-loci Y chromosome haplotype in three Italian populations". Forensic Science International.
- (2005). "Y chromosome STR polymorphisms in a Serbian population sample". Forensic Science International.
- (2006). "Population genetic study in two Transylvanian populations using forensically informative autosomal and Y-chromosomal STR markers". Forensic Science International.
- (2005). "Population data for 12 Y-chromosome STRs in a sample from Brescia (northern Italy)". Forensic Science International.
- (2003). "Y-chromosomal STR haplotypes in a Romanian population sample". International Journal of Legal Medicine.
- (2007). "Human Y-specific STR haplotypes in population of Serbia and Montenegro". Forensic Science International.
- (2009). "Population genetics of Y-chromosome STRs in a population of Northern Greeks". Forensic Science International. Genetics.
- (2006). "Y-chromosome STR haplotypes in a population sample from Switzerland (Zurich area)". Forensic Science International.
- (2008). "Evaluation of haplotype discrimination capacity of 35 Y-chromosomal short tandem repeat loci". Forensic Science International.
- (2008). "Allele frequencies and population data for 17 Y-chromosome STR loci in a Serbian population sample from Vojvodina province". Forensic Science International.
- (2004). "Population genetics of Y-chromosome STRs in a population of Podlasie, Northeastern Poland". Forensic Science International.
- (2008). "Y-chromosomal diversity in Lebanon is structured by recent historical events". American Journal of Human Genetics.
- (2007). "Y-chromosomal evidence for a limited Greek contribution to the Pathan population of Pakistan". European Journal of Human Genetics.
- (2003). "The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations". American Journal of Human Genetics.
- (2008). "Differential Y-chromosome Anatolian influences on the Greek and Cretan Neolithic". Annals of Human Genetics.
- (2004). "Y chromosome and mitochondrial DNA variation in Lithuanians". Annals of Human Genetics.
- (2009). "Population structure in contemporary Sweden--a Y-chromosomal and mitochondrial DNA analysis". Annals of Human Genetics.
- (2000). "Highly informative Y-chromosomal haplotypes by the addition of three new STRs DYS437, DYS438 and DYS439". International Journal of Legal Medicine.
- (2005). "Molecular insight into the genesis of ranked caste populations of western India based upon polymorphisms across non-recombinant and recombinant regions in genome". Genome Biology.
- (2007). "Y-chromosome genetic structure in sub-Apennine populations of Central Italy by SNP and STR analysis". International Journal of Legal Medicine.
- (2009). "Phylogeography of French male lineages". Forensic Science International.
- (October 2021). "What a difference a year makes". Italy DNA Project }}{{self-published inline.
- (28 February 2007). "Study Raises Possibility of Jewish Tie for Jefferson". The New York Times.
- (2017). "The Genetic Legacy of Zoroastrianism in Iran and India: Insights into Population Structure, Gene Flow, and Selection". The American Journal of Human Genetics.
- (2012). "Neolithic patrilineal signals indicate that the Armenian plateau was repopulated by agriculturalists". European Journal of Human Genetics.
- (2014). "Human paternal lineages, languages, and environment in the Caucasus". Human Biology.
- (2008). "Culture creates genetic structure in the Caucasus: autosomal, mitochondrial, and Y-chromosomal variation in Daghestan". BMC Genetics.
- (2001). "The Y chromosome pool of Jews as part of the genetic landscape of the Middle East". American Journal of Human Genetics.
- (2006). "Genetic Testing of Language Replacement Hypothesis in Southwest Asia". Iran and the Caucasus.
- (2011). "Y chromosome diversity among the Iranian religious groups: a reservoir of genetic variation". Annals of Human Biology.
- (2010). "The origin of Eastern European Jews revealed by autosomal, sex chromosomal and mtDNA polymorphisms". Biology Direct.
- (2009). "A Y-STR database of Iranian and Azerbaijanian minority populations". Forensic Science International. Genetics.
- (2016). "Coevolution of genes and languages and high levels of population structure among the highland populations of Daghestan". Journal of Human Genetics.
- (2012). "Paternal lineage analysis supports an Armenian rather than a Central Asian genetic origin of the Hamshenis". Human Biology.
- (2012). "Ancient migratory events in the Middle East: new clues from the Y-chromosome variation of modern Iranians". PLOS ONE.
- Yonan ''et al.'', "Y-chromosome diversity in the Assyrian Christians," (2009)
- Pavel Flegontov et al., "Genomic study of the Ket: a Paleo-Eskimo-related ethnic group with significant ancient North Eurasian ancestry," " ''Scientific Reports'' (2016)
- (2004). "The Levant versus the Horn of Africa: evidence for bidirectional corridors of human migrations". American Journal of Human Genetics.
- (2013). "Afghan Hindu Kush: where Eurasian sub-continent gene flows converge". PLOS ONE.
- (2013). "Y-chromosome and mtDNA genetics reveal significant contrasts in affinities of modern Middle Eastern populations with European and African populations". PLOS ONE.
- (2003). "Y-chromosome and mtDNA polymorphisms in Iraq, a crossroad of the early human dispersal and of post-Neolithic migrations". Molecular Phylogenetics and Evolution.
- (2009). "Geographical structure of the Y-chromosomal genetic landscape of the Levant: a coastal-inland contrast". Annals of Human Genetics.
- (1999). "Y-chromosome specific YCAII, DYS19 and YAP polymorphisms in human populations: a comparative study". Annals of Human Genetics.
- (2001). "High-resolution analysis of Y-chromosomal polymorphisms reveals signatures of population movements from Central Asia and West Asia into India". Journal of Genetics.
- (2011). "Y-chromosomal STRs in two populations from Israel and the Palestinian Authority Area: Christian and Muslim Arabs". Forensic Science International. Genetics.
- (2009). "Saudi Arabian Y-Chromosome diversity and its relationship with nearby regions". BMC Genetics.
- Marc Haber ''et al.'', "Influences of history, geography, and religion on genetic structure: the Maronites in Lebanon," ''European Journal of Human Genetics'' 2010
- (2005). "Isolates in a corridor of migrations: a high-resolution analysis of Y-chromosome variation in Jordan". Journal of Human Genetics.
- (2009). "Genetic structure of nomadic Bedouin from Kuwait". Heredity.
- (2001). "Armenian Y chromosome haplotypes reveal strong regional structure within a single ethno-national group". Human Genetics.
- (2009). "Analysis of Y-chromosomal short tandem repeat (STR) polymorphism in an Iranian Sadat population". Russian Journal of Genetics.
- (2013). "Y-chromosome variation in Tajiks and Iranians". Annals of Human Biology.
- (2017). "Turkish Cypriot paternal lineages bear an autochthonous character and closest resemblance to those from neighbouring Near Eastern populations". Annals of Human Biology.
- (2015). "Genetic profile of 17 Y-chromosome STR haplotypes in East of Iran". Forensic Science International. Genetics.
- (2006). "Iran: Tricontinental Nexus for Y-Chromosome Driven Migration". Human Heredity.
- (2011). "The coming of the Greeks to Provence and Corsica: Y-chromosome models of archaic Greek colonization of the western Mediterranean". BMC Evolutionary Biology.
- (2016). "Y-chromosome phylogeographic analysis of the Greek-Cypriot population reveals elements consistent with Neolithic and Bronze Age settlements". Investigative Genetics.
- Oleg Balanovsky ''et al.'', "Parallel Evolution of Genes and Languages in the Caucasus Region," ''Molecular Biology and Evolution'' 2011
- (2016). "Genetic structure of the Kuwaiti population revealed by paternal lineages". American Journal of Human Biology.
- (2004). "Genetic evidence concerning the origins of South and North Ossetians". Annals of Human Genetics.
- (2004). "Mitochondrial DNA and Y-chromosome variation in the caucasus". Annals of Human Genetics.
- (2005). "MtDNA and Y-chromosome variation in Kurdish groups". Annals of Human Genetics.
- (2010). "Y chromosomes of self-identified Syeds from the Indian subcontinent show evidence of elevated Arab ancestry but not of a recent common patrilineal origin". Archaeological and Anthropological Sciences.
- R. Spencer Wells ''et al.'', "The Eurasian Heartland: A continental perspective on Y-chromosome diversity," ''The National Academy of Sciences'', 2001
- (2003). "Haplotypes from the Caucasus, Turkey and Iran for nine Y-STR loci". Forensic Science International.
- (26 June 2018). "Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe". Proceedings of the National Academy of Sciences.
- (2011). "Variation in Y chromosome, mitochondrial DNA and labels of identity on Ethiopia". UCL Discovery.
- Mélanie Capredon ''et al.'', "Tracing Arab-Islamic Inheritance in Madagascar: Study of the Y-chromosome and Mitochondrial DNA in the Antemoro," ''^PLOS ONE'', 2013
- Andrea Berti ''et al.'', "YHRD Contribution," ''^YHRD'', 2016
- (2016). "Chad Genetic Diversity Reveals an African History Marked by Multiple Holocene Eurasian Migrations". American Journal of Human Genetics.
- (2009). "Y-chromosomal STR haplotypes in an Arab population from Somalia". Forensic Science International: Genetics Supplement Series.
- (2005). "Introduction of a single nucleodite polymorphism-based 'Major Y-chromosome haplogroup typing kit' suitable for predicting the geographical origin of male lineages". Electrophoresis.
- (2011). "Dissecting the within-Africa ancestry of populations of African descent in the Americas". PLOS ONE.
- (2015). "Inferring population structure and demographic history using Y-STR data from worldwide populations". Molecular Genetics and Genomics.
- (2004). "Kurdish (Iraq) and Somalian population data for 15 autosomal and 9 Y-chromosomal STR loci". International Congress Series.
- (2005). "Contrasting patterns of Y chromosome and mtDNA variation in Africa: evidence for sex-biased demographic processes". European Journal of Human Genetics.
- (2009). "A multi-perspective view of genetic variation in Cameroon". American Journal of Physical Anthropology.
- (2002). "A back migration from Asia to sub-Saharan Africa is supported by high-resolution analysis of human Y-chromosome haplotypes". American Journal of Human Genetics.
- (2005). "Y-chromosome STR haplotypes in Somalis". Forensic Science International.
- (2005). "High frequencies of Y chromosome lineages characterized by E3b1, DYS19-11, DYS392-12 in Somali males". European Journal of Human Genetics.
- (2004). "A predominantly neolithic origin for Y-chromosomal DNA variation in North Africa". American Journal of Human Genetics.
- Andreas O. Tillmar ''et al.'' "Population data of 12 Y-STR loci from a Somali population (2009)
- Ghada A. Omran ''et al.'', "Diversity of 17-locus Y-STR haplotypes in Upper (Southern) Egyptians," ''Forensic Science International: Genetics Supplement Series''2007
- Cesare de Filippo ''et al.'', "Y-chromosomal variation in Sub-Saharan Africa, insights into the history of Niger–Congo groups," ''Oxford University Press'', 2010
- (2010). "Linking the sub-Saharan and West Eurasian gene pools: maternal and paternal heritage of the Tuareg nomads from the African Sahel". European Journal of Human Genetics.
- (2007). "History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation". Molecular Biology and Evolution.
- (2007). "Tracing past human male movements in northern/eastern Africa and western Eurasia: new clues from Y-chromosomal haplogroups E-M78 and J-M12". Molecular Biology and Evolution.
- (2013). "Population genetics of 17 Y-STR markers in West Libya (Tripoli region)". Forensic Science International. Genetics.
- (2006). "Haplotypes for 13 Y-chromosomal STR loci in South Tunisian population (Sfax region)". Forensic Science International.
- (2006). "Y-chromosomal STR haplotypes in an Arab population from Libya". International Congress Series.
- Called "Wairak" and misidentified as Bantu in the studies.
- (2003). "A study of Y-chromosome microsatellite variation in sub-Saharan Africa: a comparison between F(ST) and R(ST) genetic distances". Human Biology.
- (2011). "Y-chromosomal variation in sub-Saharan Africa: insights into the history of Niger-Congo groups". Molecular Biology and Evolution.
- (2009). "Becoming Eloquent: Advances in the Emergence of Language, Human Cognition, and Modern Cultures". John Benjamins.
- (2004). "AmpFℓSTR Identifiler STR Allele Frequencies and PowerPlex Y-STR Haplotype Frequencies of the Meru Population of Northern Tanzania". University of Nevada.
- (2010). "Little genetic differentiation as assessed by uniparental markers in the presence of substantial language variation in peoples of the Cross River region of Nigeria". BMC Evolutionary Biology.
- (2003). "Y-chromosome lineages in Cabo Verde Islands witness the diverse geographic origin of its first male settlers". Human Genetics.
- (2011). "Y-STR haplotypes in three ethnic linguistic groups of Angola population". Forensic Science International. Genetics.
- (2012). "Population genetic data for 17 Y STR markers from Benghazi (East Libya)". Forensic Science International. Genetics.
- (2015). "Genetic population study of Y-chromosome markers in Benin and Ivory Coast ethnic groups". Forensic Science International. Genetics.
- (2010). "Digging deeper into East African human Y chromosome lineages". Human Genetics.
- (2002). "Y-chromosome analysis in Egypt suggests a genetic regional continuity in Northeastern Africa". Human Biology.
- (2016). "Palenque de San Basilio in Colombia: genetic data support an oral history of a paternal ancestry in Congo". Proceedings: Biological Sciences.
- (2000). "Y chromosome STR haplotypes in four populations from northwest Africa". International Journal of Legal Medicine.
- (2015). "Genetic diversity and haplotype structure of 21 Y-STRs, including nine noncore loci, in South Tunisian Population: Forensic relevance". Electrophoresis.
- (2015). "Sousse: extreme genetic heterogeneity in North Africa". Journal of Human Genetics.
- (2005). "Y-chromosomal STR haplotypes in three ethnic groups and one cosmopolitan population from Tunisia". Forensic Science International.
- (2016). "Group membership, geography and shared ancestry: Genetic variation in the Basotho of Lesotho". American Journal of Physical Anthropology.
- (2011). "Allele frequencies and population data for 17 Y-STR loci (The AmpFlSTR Y-filer) in Casablanca resident population". Forensic Science International. Genetics.
- (2016). "Refining the Y chromosome phylogeny with southern African sequences". Human Genetics.
- (2016). "The paternal ancestry of Uttarakhand does not imitate the classical caste system of India". Journal of Human Genetics.
- (2011). "Indian Siddis: African Descendants with Indian Admixture". The American Journal of Human Genetics.
- (2007). "Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations". BMC Evolutionary Biology.
- (2006). "Polarity and temporality of high-resolution y-chromosome distributions in India identify both indigenous and exogenous expansions and reveal minor genetic influence of Central Asian pastoralists". American Journal of Human Genetics.
- (2012). "Genetic affinities of the central Indian tribal populations". PLOS ONE.
- (2012). "Population differentiation of southern Indian male lineages correlates with agricultural expansions predating the caste system". PLOS ONE.
- (2015). "Population genetics of 17 Y-chromosomal STRs loci in Garo and Santal tribal populations in Bangladesh". International Journal of Legal Medicine.
- (2007). "Y-chromosomal STR haplotypes in two endogamous tribal populations of Karnataka, India". Journal of Forensic Sciences.
- (2001). "Y-chromosome SNP haplotypes suggest evidence of gene flow among caste, tribe, and the migrant Siddi populations of Andhra Pradesh, South India". European Journal of Human Genetics.
- R. Cordaux et al. "Independent Origins of Indian Caste and Tribal Paternal Lineages"
- (2006). "Genetic affinities among the lower castes and tribal groups of India: inference from Y chromosome and mitochondrial DNA". BMC Genetics.
- (2009). "The Indian origin of paternal haplogroup R1a1* substantiates the autochthonous origin of Brahmins and the caste system". Journal of Human Genetics.
- (August 2004). "The northeast Indian passageway: a barrier or corridor for human migrations?". Mol. Biol. Evol..
- (2001). "Ethnic populations of India as seen from an evolutionary perspective". Journal of Biosciences.
- А. Х. Маргулана, "Первые результаты работы Лаборатории популяционной генетики," "ИНСТИТУТ ОБЩЕЙ ГЕНЕТИКИ И ЦИТОЛОГИИ" (2016), http://iggc.kz/wp-content/uploads/2017/03/Rezultaty-raboty-Lab-Pop-Gen-noyab-2016.pdf {{Webarchive. link. (17 January 2021)
- (2002). "A human genome diversity cell line panel". Science.
- (2010). "Y-chromosome distributions among populations in Northwest China identify significant contribution from Central Asian pastoralists and lesser influence of western Eurasians". Journal of Human Genetics.
- (2006). "Male demography in East Asia: a north-south contrast in human population expansion times". Genetics.
- (2015). "Mitochondrial DNA and the Y chromosome suggest the settlement of Madagascar by Indonesian sea nomad populations". BMC Genomics.
- (2013). "Genetic evidence of paleolithic colonization and neolithic expansion of modern humans on the tibetan plateau". Molecular Biology and Evolution.
- (2004). "The northeast Indian passageway: a barrier or corridor for human migrations?". Molecular Biology and Evolution.
- (2012). "Afghanistan's Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events". PLOS ONE.
- (2015). "Genetic evidence of a recent Tibetan ancestry to Sherpas in the Himalayan region". Scientific Reports.
- (2015). "Y-chromosome descent clusters and male differential reproductive success: young lineage expansions dominate Asian pastoral nomadic populations". European Journal of Human Genetics.
- (2011). "Extended Y chromosome investigation suggests postglacial migrations of modern humans into East Asia via the northern route". Molecular Biology and Evolution.
- М.К. Жабагин et al., "The relation between the Y-chromosomal variation and the clan structure: the gene pool of the steppe aristocracy and the steppe clergy of the Kazakhs," "Russian Journal of Genetics," (2014),
- (2015). "Haplotype analysis of the polymorphic 40 Y-STR markers in Chinese populations". Forensic Science International. Genetics.
- Niaz M. Achakzai ''et al.'', "Y-chromosomal STR analysis in the Pashtun population of Southern Afghanistan," ''Molecular Biology and Evolution'' 2011
- Sadia Tabassum ''et al.'', "A comprehensive Y-STR portrait of Yousafzai's population," ''International Journal of Legal Medicine'' 2017
- (2011). "Polymorphisms of Y-STRs in Uygur and Kazak Ethnic in Xinjiang". Xinjiang Medical University.
- (2016). "Genetic analysis of 17 Y-STR loci in Han and Korean populations from Jilin Province, Northeast China". Forensic Science International. Genetics.
- (2016). "Population genetics of 23 Y-STR loci in the Mongolian minority population in Inner Mongolia of China". International Journal of Legal Medicine.
- (2010). "Genetic polymorphism of 11 Y-chromosomal STR loci in Yunnan Han Chinese". Forensic Science International. Genetics.
- (2008). "Y-chromosomal STRs haplotypes in Chinese Hui ethnic group samples". Forensic Science International. Genetics.
- (2009). "Haplotype diversity of 17 Y-chromosomal STRs in three native Sarawak populations (Iban, Bidayuh and Melanau) in East Malaysia". Forensic Science International. Genetics.
- (2007). "The genetic male legacy from El Salvador". Forensic Science International.
- (2009). "Y-chromosome haplotypes defined by 17 STRs included in AmpFlSTR Yfiler PCR Amplification Kit in a multi ethnical population from El Beni Department (North Bolivia)". Legal Medicine (Tokyo, Japan).
- (2010). "Genetic make up and structure of Colombian populations by means of uniparental and biparental DNA markers". American Journal of Physical Anthropology.
- (2012). "CoanCestría de apellidos y linajes del. Cromosoma y en el noroeste de Colombia: una herramienta útil para establecer migración entre poblaciones". Revista Colombiana de Antropología.
- (2016). "A comprehensive Y-STR portrait of Argentinean populations". Forensic Science International. Genetics.
- (2008). "Y-chromosome STR haplotypes in three different population groups from Ecuador (South America)". Journal of Forensic Sciences.
- (2014). "Between Andes and Amazon: the genetic profile of the Arawak-speaking Yanesha". American Journal of Physical Anthropology.
- Luz Angela Alonso Morales. (2013). "Caracterización de la población humana de los departamentos de Tolima y Huila. Perspectivas: demográficas, genéticas y socioculturales". Universidad Nacional de Colombia.
- (2011). "Paternal lineages signal distinct genetic contributions from British Loyalists and continental Africans among different Bahamian islands". American Journal of Physical Anthropology.
- (2009). "Comparison of Y-chromosome haplogroup frequencies in eight Provinces of Argentina". Forensic Science International: Genetics Supplement Series.
- (2011). "Male lineages in South American native groups: evidence of M19 traveling south". American Journal of Physical Anthropology.
- (2008). "Y chromosome microsatellite genetic variation in two Native American populations from Argentina: population stratification and mutation data". Forensic Science International. Genetics.
- (2015). "Exploring the Y Chromosomal Ancestry of Modern Panamanians". PLOS ONE.
- (2010). "Y-chromosome lineages in native South American population". Forensic Science International: Genetics.
- (2000). "Strong Amerind/white sex bias and a possible Sephardic contribution among the founders of a population in northwest Colombia". American Journal of Human Genetics.
- (2010). "Inferring continental ancestry of argentineans from Autosomal, Y-chromosomal and mitochondrial DNA". Annals of Human Genetics.
- (2012). "Distribution of Y chromosomal STRs loci in Mayan and Mestizo populations from Guatemala". Forensic Science International. Genetics.
- (2016). "Analysis of Y-chromosome STRs in Chile confirms an extensive introgression of European male lineages in urban populations". Forensic Science International. Genetics.
- (2015). "Male-specific contributions to the Brazilian population of Espirito Santo". International Journal of Legal Medicine.
- (2007). "Population data for 15 Y-chromosome STRs in a population sample from Quito (Ecuador)". Forensic Science International.
- (2016). "MtDNA and Y-chromosomal diversity in the Chachapoya, a population from the northeast Peruvian Andes-Amazon divide". American Journal of Human Biology.
- (2010). "Reconstructing the population history of Nicaragua by means of mtDNA, Y-chromosome STRs, and autosomal STR markers". American Journal of Physical Anthropology.
- (2015). "Admixture and Genetic Diversity Distribution Patterns of Non-Recombining Lineages of Native American Ancestry in Colombian Populations". PLOS ONE.
- (2011). "mtDNA and Y-chromosome diversity in Aymaras and Quechuas from Bolivia: different stories and special genetic traits of the Andean Altiplano populations". American Journal of Physical Anthropology.
- (2014). "Genetic Analysis of 17 Y-STRs in a Mestizo Population from the Central Valley of Mexico". Human Biology.
- (2013). "Colombia's racial crucible: Y chromosome evidence from six admixed communities in the Department of Bolivar". Annals of Human Biology.
- (2010). "Population data for 12 Y-chromosome STR loci in a sample from El Salvador". Legal Medicine.
- (2012). "Y-chromosomal diversity in Haiti and Jamaica: contrasting levels of sex-biased gene flow". American Journal of Physical Anthropology.
- (2007). "Y chromosome STR haplotypes in the Caribbean city of Cartagena (Colombia)". Forensic Science International.
- (2008). "Haplotype data for 12 Y-chromosome STR loci from Costa Rica". Forensic Science International: Genetics Supplement Series.
- (2010). "Y-chromosomal STR haplotype diversity in males from Santa Catarina, Brazil". Journal of Forensic and Legal Medicine.
- (2007). "Mitochondrial and Y chromosome diversity in the English-speaking Caribbean". Annals of Human Genetics.
- (2009). "Population data for 12 Y-chromosome STR loci in a sample from Honduras". Legal Medicine.
- (2012). "Haplotype diversity of 17 Y-str loci in an admixed population from the Brazilian Amazon". Genetics and Molecular Biology.
- (2015). "Allele frequencies for 15 autosomal STR loci and haplotype data for 17 Y-STR loci in a population from Belize". International Journal of Legal Medicine.
- (2016). "Male-specific contributions to the Brazilian population of Espirito Santo". International Journal of Legal Medicine.
- (20 August 2018). "Ancient DNA from Chalcolithic Israel reveals the role of population mixture in cultural transformation". Nature Communications.
- [https://www.jpost.com/Israel-News/DNA-study-in-Israeli-cave-sheds-light-on-origins-of-Chalcolithic-culture-565293 DNA STUDY IN ISRAELI CAVE SHEDS LIGHT ON ORIGINS OF CHALCOLITHIC CULTURE]
- [https://www.timesofisrael.com/anomalous-blue-eyed-people-came-to-israel-6500-years-ago-from-iran-dna-shows/ Anomalous blue-eyed people came to Israel 6,500 years ago from Iran, DNA shows]
- "T-L162 YTree".
- (2020). "Genomic History of Neolithic to Bronze Age Anatolia, Northern Levant, and Southern Caucasus". Cell.
- "T-CTS11451 YTree".
- (2002). "Ethiopians and Khoisan Share the Deepest Clades of the Human Y-Chromosome Phylogeny". The American Journal of Human Genetics.
- (2004). "Y chromosome haplogroups of elite Ethiopian endurance runners". Human Genetics.
- (April 2007). "Thomas Jefferson's Y chromosome belongs to a rare European lineage". Am. J. Phys. Anthropol..
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Haplogroup T-M184 — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report