From Surf Wiki (app.surf) — the open knowledge base
Haplogroup NO1
Human Y-chromosome DNA haplogroup
Human Y-chromosome DNA haplogroup
| Field | Value |
|---|---|
| name | NO1 |
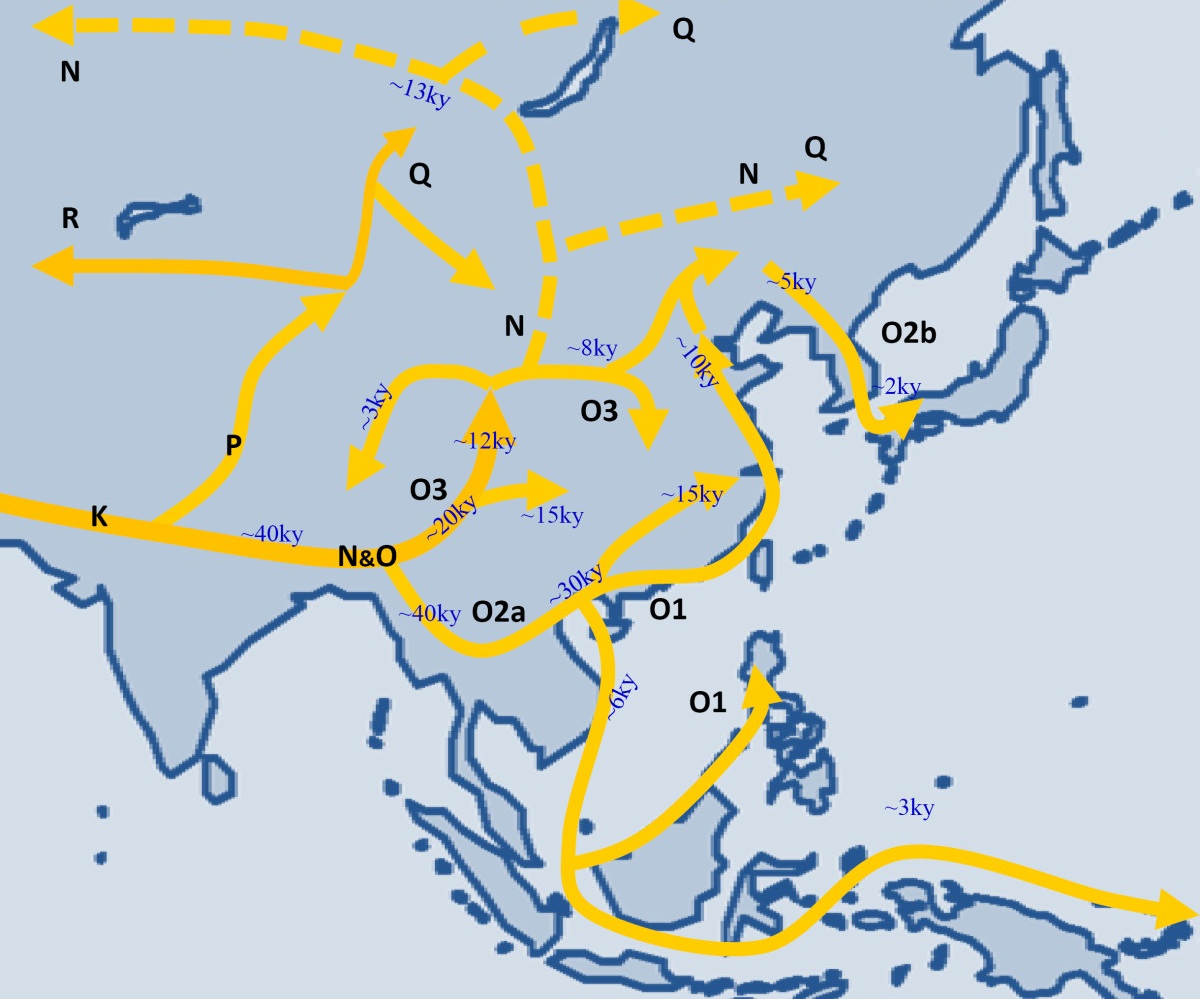
| map | File:Haplogroup NO.png |
| origin-date | Formed circa 45,400–45,840 years BP |
| (NO/NO1, based on YFull 2017, and previous estimates of: 41,500 [95% CI 37,400 45,600] years BP, 48,871 [95% CI 37,095 58,831] years BP, and 50,800 or 43,500 years BP) | |
| origin-place | Southeast Asia or East Asia (Northern China) |
| TMRCA | Circa 41,500 – 40,067 years BP (NO/NO1, based on YFull 2017, and previous estimates of: 36,800 [95% CI 34,300 39,300] years BP, 41,900 [31,294 51,202] years BP, and 44,700 or 38,300 years BP depending on mutation rate) |
| ancestor | K-M2313 (M2313/Z4858) |
| descendants | N (M231) and O (M175). |
| mutations | M214/Page39; F176/M2314; CTS5858/M2325/F346; CTS11572 |
| origin-date = Formed circa 45,400–45,840 years BP (NO/NO1, based on YFull 2017, and previous estimates of: 41,500 [95% CI 37,400 45,600] years BP, 48,871 [95% CI 37,095 58,831] years BP, and 50,800 or 43,500 years BP) | origin-place = Southeast Asia or East Asia (Northern China)
The location of NO1 at the SNP M214 follows the taxonomy set out by Karmin et al. 2022, and conforms to a structure shown by ISOGG (2022). (However neither Karmin nor ISOGG has integrated one of the findings of Poznik et al. 2016: that there was a subclade generation both above haplogroup NO1 (M214) and beneath K-M2308. That is, both ISOGG and Karmin et al. continued to equate K2a with haplogroup NO.)
Before 2016, the subclades compromising both NO and NO1 were not recognised, and were regarded as synonymous with K2a. Researchers such as David Poznik (Poznik et al. 2016) documented examples of previously unknown subclades of haplogroup K2, in both ancient remains and living individuals, which: (firstly) had several, varying suites of the SNPs regarded previously as uniquely defining K2a and NO, but also (secondly) lacked any of the SNPs specifically identifying haplogroups Haplogroup N (M231) and Haplogroup O (M175). This demonstrated conclusively that multiple stages of development separated K2a from NO, which therefore constituted "grandparent" and "grandchild" clades. Poznik et al. 2016 used the name "K2a1" (a.k.a. K-M2313) for the Y-DNA of some of the individuals who belonged to K2a(xK2a*,NO), while also mentioning that K-M2313 did not include all examples of K2a(xK2a*,NO).
As of 2022, the International Society of Genetic Genealogy (ISOGG) refers to NO-M214 as "NO1", and to K2a (M2308)/K-M2313 as "NO". There may be at least one other primary branch of NO: the ISOGG official Y-DNA haplogroup tree lists a haplogroup known as "NO1~" (CTS707/M2306) alongside NO-M214 (which ISOGG refers to as "NO1"). The tilde () indicates that its exact position of NO1 in the phylogeny is unknown. It may be a primary branch or sibling of NO, it may be a primary branch or sibling of K2a, K-M2313, or it may instead be a primary branch of K2a.
Based on the projected origins of K2a, K-M2313, and the basal haplogroups N* and O* respectively, NO* probably originated in East Asia.
Distribution

While there is some evidence of NO* being found in living individuals, these examples are not well-researched. Further research may instead identify them as belonging to N* (M231), N1, or the provisional subclade N2 (F3373/M2283/Page56/S323). These cases include:
- 5.7% (2/35) of Bouyei males, in China or Vietnam;
- a pool of four samples of Japanese males at 2.9% (6/210), particularly in Tokushima Prefecture at a rate of 5.7% (4/70), and;
- small proportions of samples from Yizu, Malays, Sô, Mongolians, Daurs, Manchurian Evenks, Hezhes, Huis, Yaos, South Koreans, Fiji (1/107 = 0.9%), Futuna 5%, Niue 3.5%, Tuvalu 3.6%, and Samoa 3%.
Members of Haplogroup NO* include a Telugu of Indian origin sampled in the United Kingdom and a Malay sampled in Singapore.
Two sets of ancient remains previously considered as possibly belonging to NO have since been reclassified upstream to K2a.
- Ust'-Ishim man dates from approximately 45,000 BP and was found in Omsk Oblast, Russia. (Until 2016 these remains were erroneously classified as K2*.)
- Oase 1: the remains found in Romania of a male who lived 37,000-42,000 years BP.
Likewise, cases previously regarded as possible examples of NO* or NO1*, and since ruled out, include:
- two Han Chinese males previously found to be negative for M175 (i.e. Haplogroup O) and LLY22g (an obsolete, possibly inaccurate marker for N1), have subsequently have been found to belong to N* (N-M231), and;
- a clade first identified in South India, defined by the SNP M147 and labelled "pre-NO" and "Haplogroup X", among other names, was found to be a sibling of NO (K2a) within Haplogroup K2 (K-M526); the new clade was renamed K2e. A 2024 study showed that several remains from Ilsenhöhle in Ranis, Germany had haplogroup NO like the Ust'Ishim remains. The Ranis individuals were more related to the Zlatý Kůň individuals and did not contribute to later Out of Africans, similar to Ust'Ishim and Oase 1 individuals. However, there is evidence of some "genetic continuity and persistence in Eurasia" due to most extant East Asian paternal lineages belonging to the downstream NO. Haplogroups O-M175 and N-M231 were introduced to East Asia via the southern route although their exact origins are unclear. Haplogroup NO1 is also observed in the modern inhabitants of Tak province from Thailand (14.23%). Its presence is attributed to complex gene flow from multiple sources such as Karen, Lawa and Khuen.
Subclades
Phylogenetic tree
This phylogeny of haplogroups K2a, K2a1, and NO is based on YFull 2018, Poznik 2016, ISOGG 2018, Karafet 2008.
K2a K-M2308 (M2308) Found only in the ancient remains "Ust'-Ishim man" (c. 45,000 BP) and "Oase 1" (c. 39,500 BP).
- K2a1 K-M2313 (M2313/Z4858) Named by Poznik 2016; previously not distinguished from K2a.
- NO K-M214 (M214/Page39; F176/M2314; CTS5858/M2325/F346; CTS11572) Poznik 2016 and ISOGG 2018 distinguish between NO1 (M214), N and O.
- N M231; CTS2947/M2175; Z4891; CTS10118
- O M175/P186/P191/P196; F369/M1755; F380/M1757/S27659
- NO K-M214 (M214/Page39; F176/M2314; CTS5858/M2325/F346; CTS11572) Poznik 2016 and ISOGG 2018 distinguish between NO1 (M214), N and O.
The position of NO1~ (CTS707/M2306), a subclade of K2a1 or NO, in this phylogeny is unclear.
References
References
- [https://www.yfull.com/tree/K-M2335/ ''YFull YTree v5.08'', 2017, "K-M2335"] (9 December 2017); [http://www.phylotree.org/Y/marker_list.htm ''PhyloTree'', 2017, "Details of the Y-SNP markers included in the minimal Y tree"] (9 December 2017); [https://www.genetichomeland.com/welcome/dnamarkerindex.asp ''GeneticHomeland.com'', 2016, DNA Marker Index Chromosome Y V4208] (9 December 2017).
- [https://www.yfull.com/tree/NO/ YFull] Haplogroup YTree v5.06 at 25 September 2017
- (April 2015). "A recent bottleneck of Y chromosome diversity coincides with a global change in culture". Genome Research.
- (2007). "A counter-clockwise northern route of the Y-chromosome haplogroup N from Southeast Asia towards Europe". European Journal of Human Genetics.
- 『DNA・考古・言語の学際研究が示す新・日本列島史 日本人集団・日本語の成立史』
- ISOGG, [https://isogg.org/tree/ISOGG_YDNATreeTrunk.html ''Y-DNA Haplogroup Tree 2018''] (17 January 2018).
- (June 2016). "Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences". Nature Genetics.
- (2 March 2022). "Episodes of Diversification and Isolation in Island Southeast Asian and Near Oceanian Male Lineages". Molecular Biology and Evolution.
- (2013). "Inferring human history in East Asia from Y chromosomes". Investigative Genetics.
- (April 2006). "Male Demography in East Asia: A North–South Contrast in Human Population Expansion Times". Genetics.
- (January 2006). "Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes". Journal of Human Genetics.
- (2014). "Genome sequence of a 45,000-year-old modern human from western Siberia". Nature.
- (August 2010). "Major East-West Division Underlies Y Chromosome Stratification across Indonesia". Molecular Biology and Evolution.
- (20 February 2025). "Earliest modern human genomes constrain timing of Neanderthal admixture". Nature.
- (May 2024). "Evolutionary profiles and complex admixture landscape in East Asia: New insights from modern and ancient Y chromosome variation perspectives". Heliyon.
- Jaisamut, Kitipong. (2025). "Paternal genetic landscape of contemporary Thai populations in the borderland provinces of Thailand and Myanmar". Scientific Reports.
- (May 2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Haplogroup NO1 — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report